tidyr::nest() and purrr:map() can be used to
easily fit the same model to different subsets of a single dataframe.
There are many
tutorials
available to help guide you
through this process. There are substantially fewer (none I’ve been able
to find) that show you how to use these two functions to fit the same
model to different features from your dataframe.
While the former involves splitting your data into different subsets by row, the latter involves cycling through different columns. I recently confronted a problem where I had to run many models, including just one predictor at a time from large pool of candidate predictors, while also including a standard set of control variables in each.1 Given the (apparent) absence of tutorials on fitting the same model to different features from a dataframe using these functions, I decided to write up the solution I reached in the hope it might be helpful to someone else.2 Start by loading the following packages:
library(tidyverse)
library(broom)
library(modelsummary)
library(kableExtra)
library(nationalparkcolors)
We’ll start with a recap of the subsetting approach, then build on it to
cycle through features instead of subsets of the data. This code is
similar to the official tidyverse
tutorial above, but
pipes the output directly to a ggplot() call to visualize the results.
mtcars %>%
nest(data = -cyl) %>% # split data by cylinders
mutate(mod = map(data, ~lm(mpg ~ disp + wt + am + gear, data = .x)),
out = map(mod, ~tidy(.x, conf.int = T))) %>% # tidy model to get coefs
unnest(out) %>% # unnest to access coefs
mutate(sig = sign(conf.low) == sign(conf.high), # p <= .05
cyl = as.factor(cyl)) %>% # factor for nicer plotting
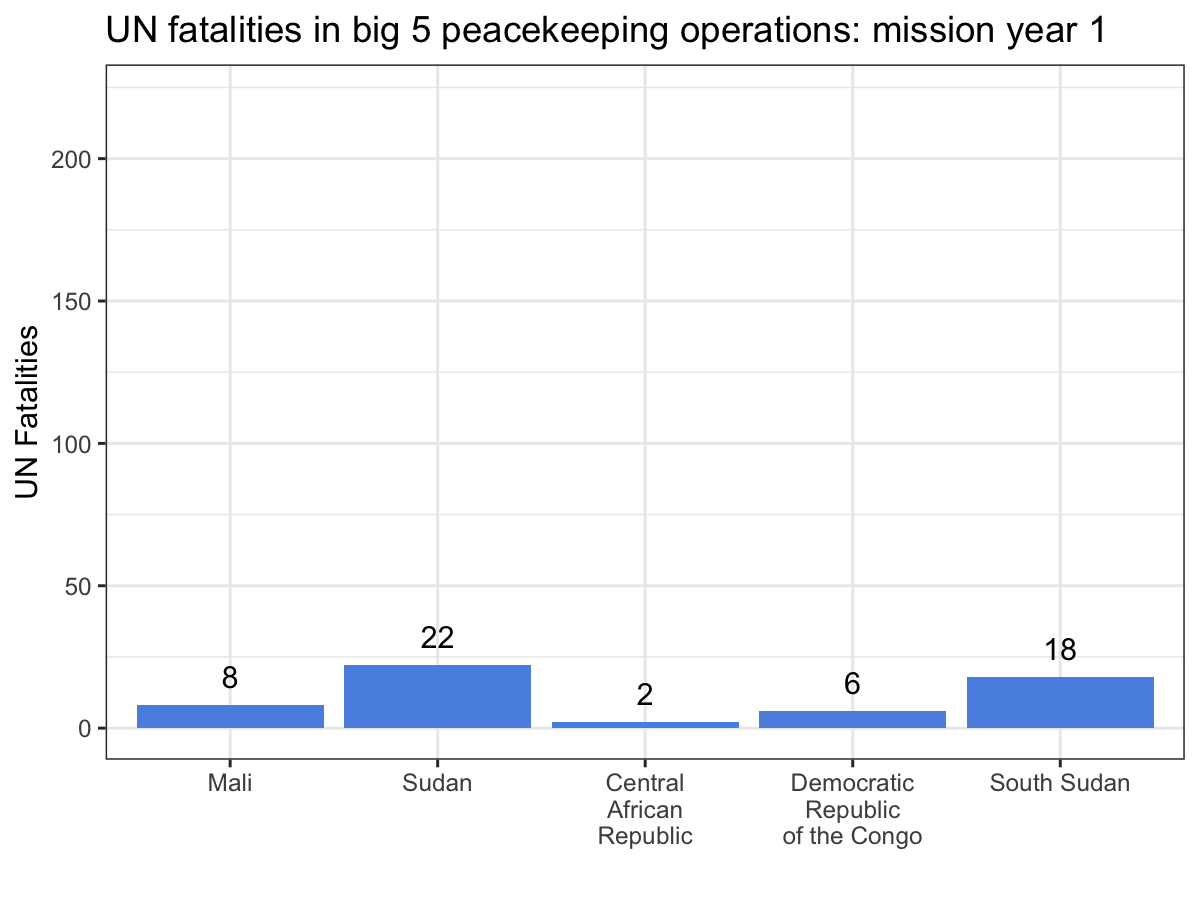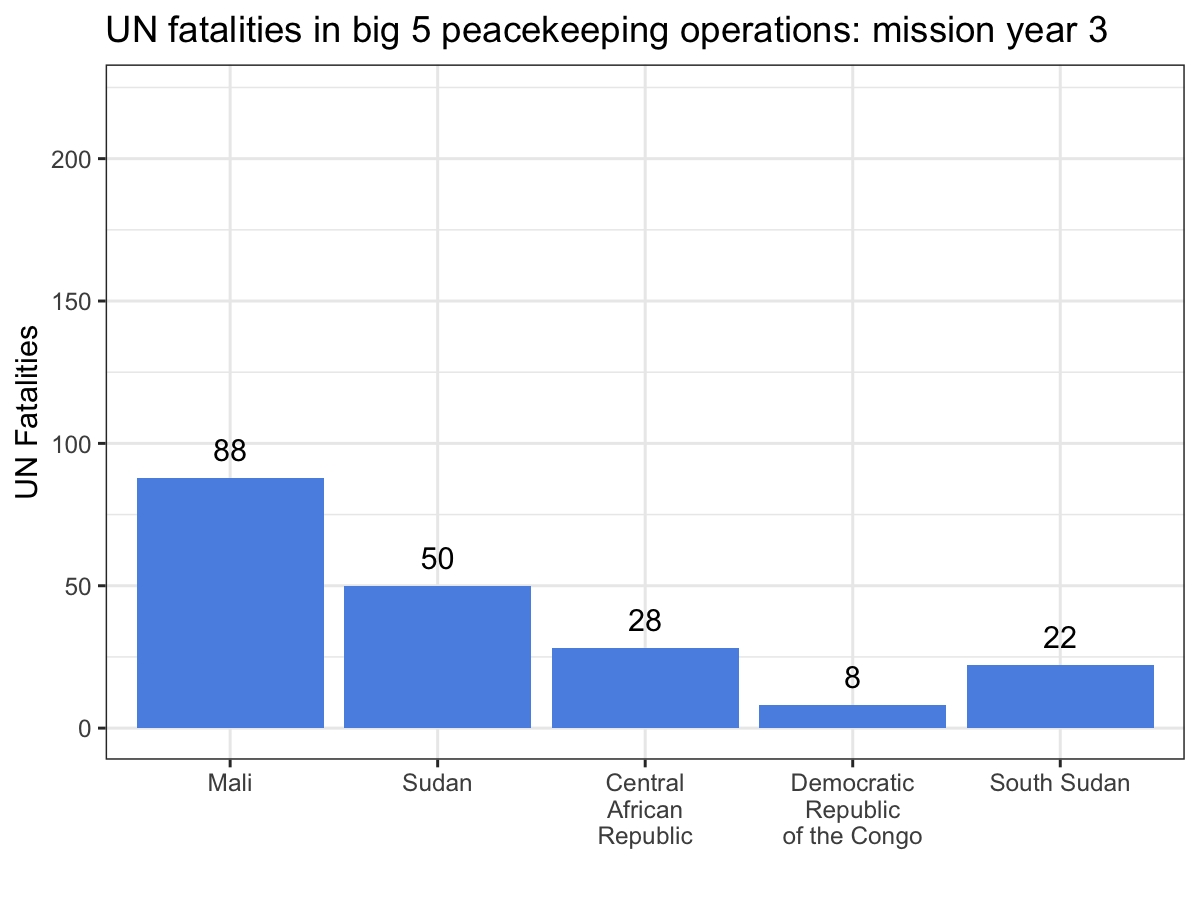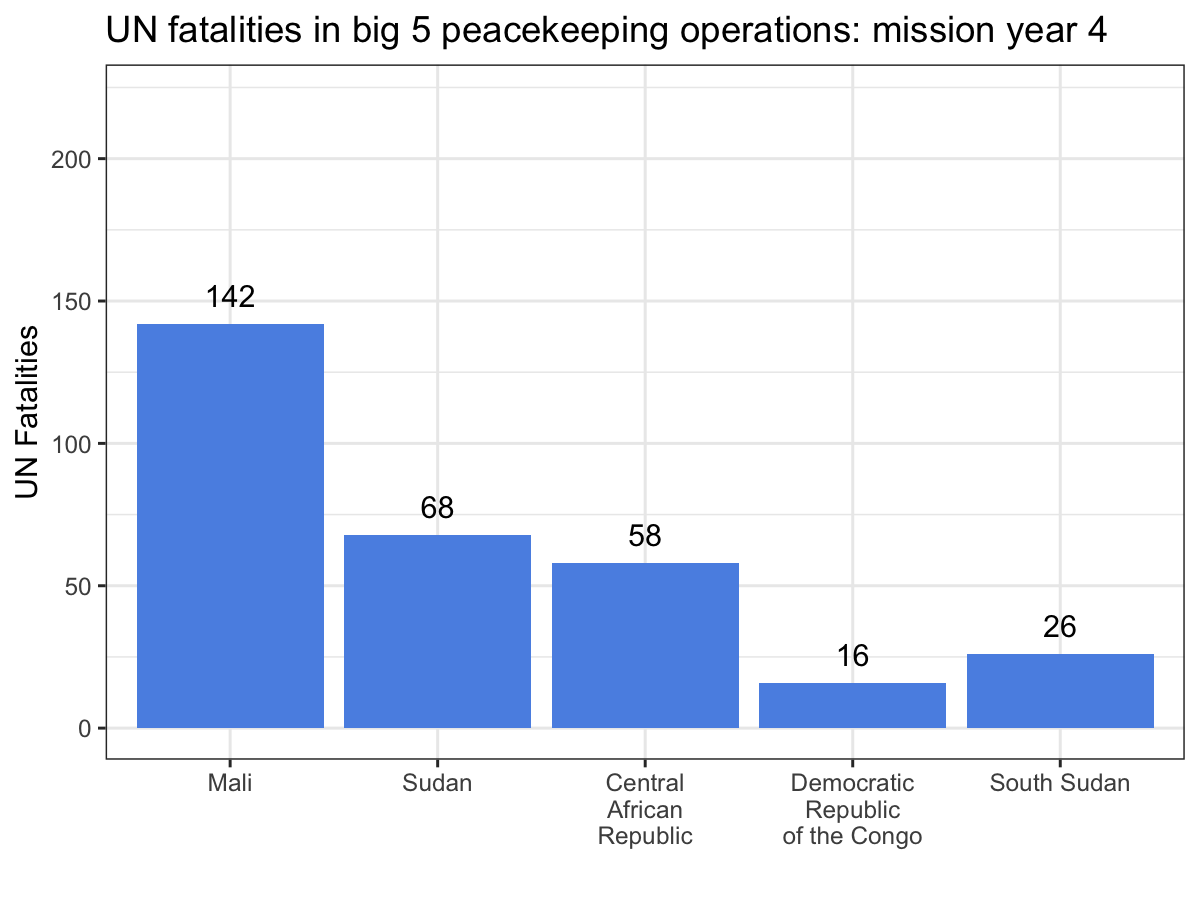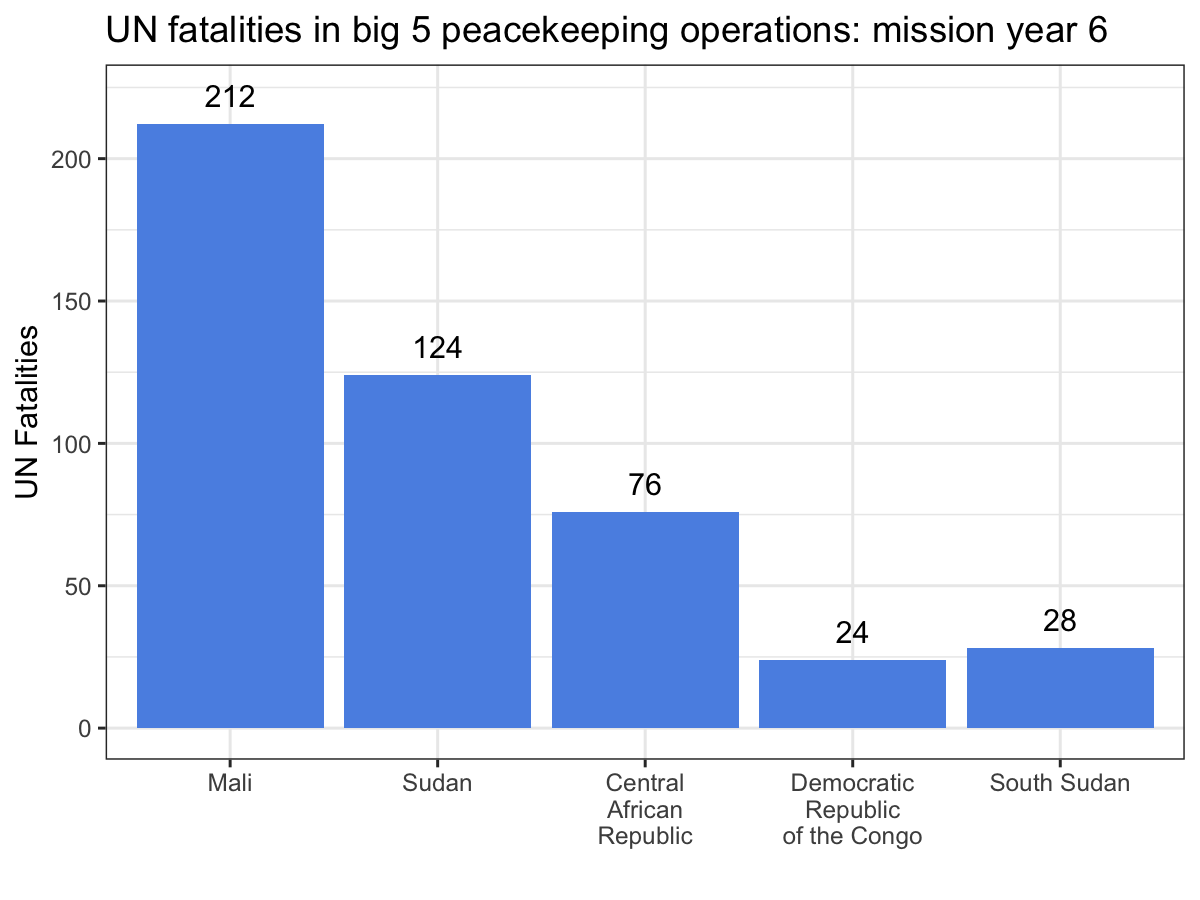
filter(term == 'disp') %>%
ggplot(aes(x = cyl, y = estimate, ymin = conf.low, ymax = conf.high,
color = sig)) +
geom_pointrange() +
geom_hline(yintercept = 0, lty = 2, color = 'grey60') +
scale_color_manual(name = 'Statistical significance',
labels = str_to_title,
values = park_palette('Saguaro')) +
labs(x = 'Cylinders', y = "Coefficient estimate") +
theme_bw() +
theme(legend.position = 'bottom')
Multiple predictors
The first thing we have to do is create a custom fuction because we now
need to be able to specify different predictors in different runs of the
model. The code below is very similar to the code above, except that
we’re defining the formula in lm() via the formula() function, which
parses a character object that we’ve assembled via str_c(). The net
effect of this is to fit a model where the pred argmument to
func_var() is the first predictor. This lets us use an external
function to supply different values to pred. Then we use
broom::tidy() to create a tidy dataframe of point estimates and
measures of uncertainty from the model and store them in a variable
called out. Finally, mutate(pred = pred) creates a variable named
pred in the output dataframe that records what the predictor used to
fit the model was. We could retrieve this from the mod list-column,
but this is approach is simpler both to extract the predictor
programtically and to visually inspect the data. We use then
purr::map_dfr() to generate a dataframe where each row corresponds to
a model with with a different predictor.
func_var <- function(pred, dataset) {
dataset %>%
nest(data = everything()) %>%
mutate(mod = map(data, ~lm(formula(str_c('mpg ~ ' , pred, # substitute pred
' + wt + am + gear')),
data = .x)),
out = map(mod, ~tidy(.x, conf.int = T))) %>%
mutate(pred = pred) %>%
return()
}
## predictors of interest
preds <- c('disp', 'hp', 'drat')
## fit models with different predictors
mods_var <- map_dfr(preds, function(x) func_var(x, mtcars))
## inspect
mods_var
## # A tibble: 3 × 4
## data mod out pred
## <list> <list> <list> <chr>
## 1 <tibble [32 × 11]> <lm> <tibble [5 × 7]> disp
## 2 <tibble [32 × 11]> <lm> <tibble [5 × 7]> hp
## 3 <tibble [32 × 11]> <lm> <tibble [5 × 7]> drat
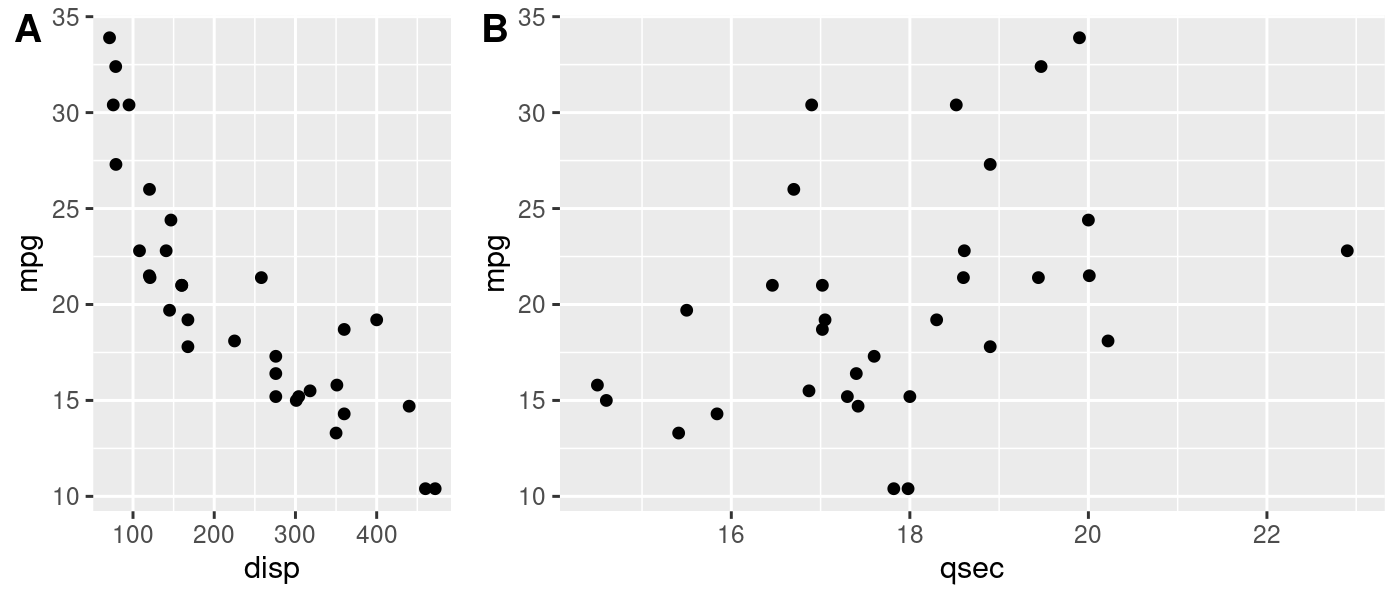
Plots
You can see our original dataframe that we condensed down into data
with nest(), the model object in mod, the tidied model output in
out, and finally the predictor used to fit the model in pred. Using
unnest(), we can unnest the out object and get a dataframe we can
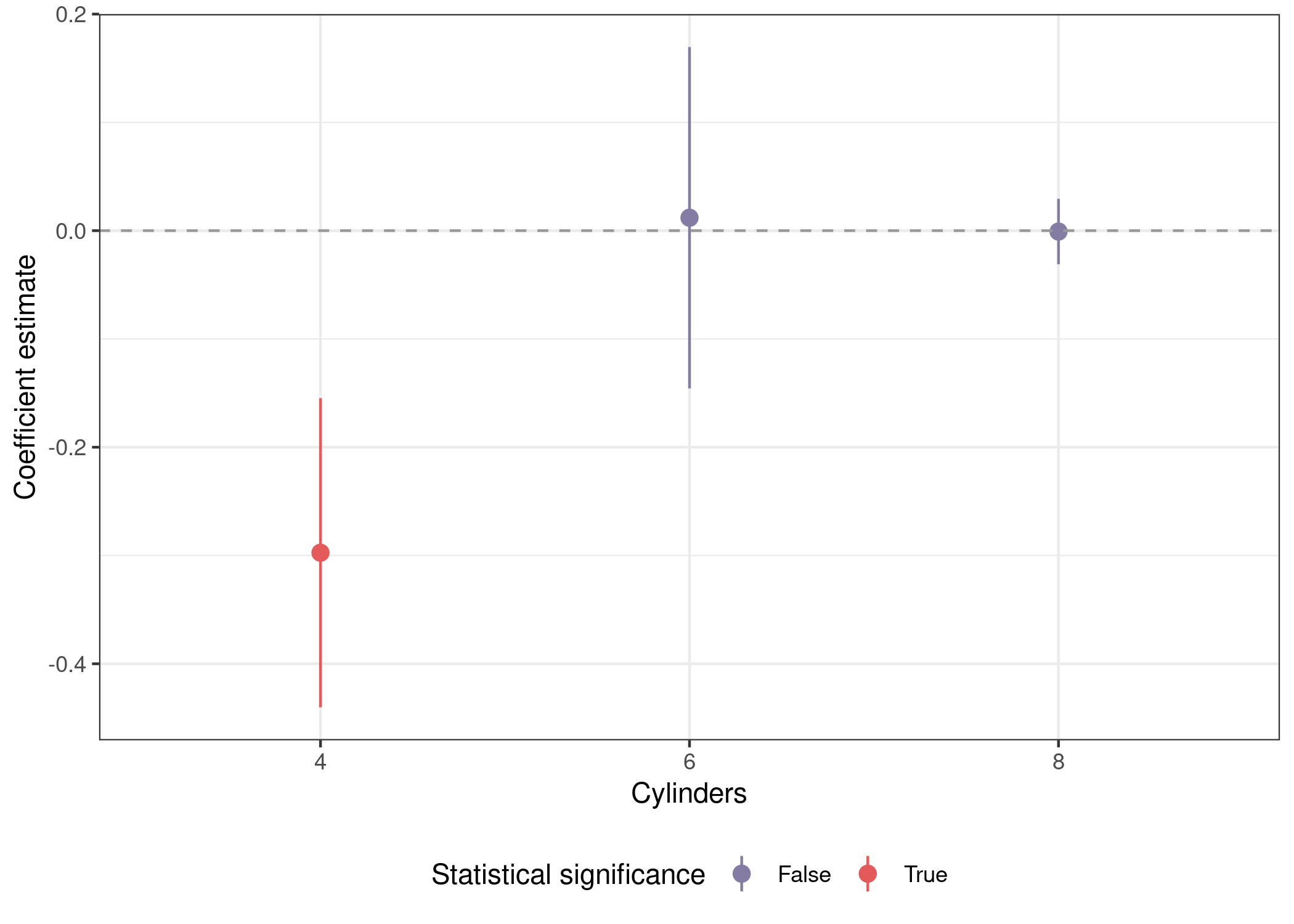
use to plot the main coefficient estimate from each of our three models.
mods_var %>%
unnest(out) %>%
mutate(sig = sign(conf.low) == sign(conf.high)) %>%
filter(term %in% preds) %>%
ggplot(aes(x = term, y = estimate, ymin = conf.low, ymax = conf.high,
color = sig)) +
geom_pointrange() +
geom_hline(yintercept = 0, lty = 2, color = 'grey60') +
scale_color_manual(name = 'Statistical significance',
labels = str_to_title,
values = park_palette('Saguaro')) +
labs(x = 'Predictor', y = "Coefficient estimate") +
theme_bw() +
theme(legend.position = 'bottom')
Tables
Things get slightly more complicated when we want to represent our
results textually instead of visually. We can use the excellent
modelsummary::modelsummary() function to create our table, but we need
to supply a list of model objects, rather than the unnested dataframe we
created above to plot the results. We can use the split() function to
turn our dataframe into a list, and by using split(seq(nrow(.))),
we’ll create one list item for each row in our dataframe.
Since each list item will be a one row dataframe, we can use lapply()
to cycle through the list. The mod object in each one row dataframe is
itself a list-column, so we need to index it with [[1]] to properly
access the model object itself.3 The last step is a call to
unname(), which will drop the automatically generated list item names
of 1, 2, and 3, allowing modelsummary() to use the default names
for each model column in the output.
tab_coef_map = c('disp' = 'Displacement', # format coefficient labels
'hp' = 'Horsepower',
'drat' = 'Drive ratio',
'wt' = 'Weight (1000 lbs)',
'am' = 'Manual',
'gear' = 'Gears',
'(Intercept)' = '(Intercept)')
mods_var %>%
split(seq(nrow(.))) %>% # list where each object is a one row dataframe
lapply(function(x) x$mod[[1]]) %>% # extract model from data dataframe
unname() %>% # remove names for default names in table
modelsummary(coef_map = tab_coef_map, stars = c('*' = .05))
Bonus
Now, let’s combine both approaches. We’re going to be splitting our
dataframe into three sub-datasets by number of cylinders while also
fitting the same model three times with 'disp', 'hp', and 'drat'
as predictors. The only changes to func_var() are to omit cyl from
the nesting, and to recode it as a factor to treat it as discrete axis
labels.
func_var_obs <- function(pred, dataset) {
dataset %>%
nest(data = -cyl) %>%
mutate(mod = map(data, ~lm(formula(str_c('mpg ~ ' , pred,
' + wt + am + gear')),
data = .x)),
out = map(mod, ~tidy(.x, conf.int = T)),
cyl = as.factor(cyl),
pred = pred) %>%
select(-data) %>%
return()
}
preds <- c('disp', 'hp', 'drat')
mods_var_obs <- map_dfr(preds, function(x) func_var_obs(x, mtcars))
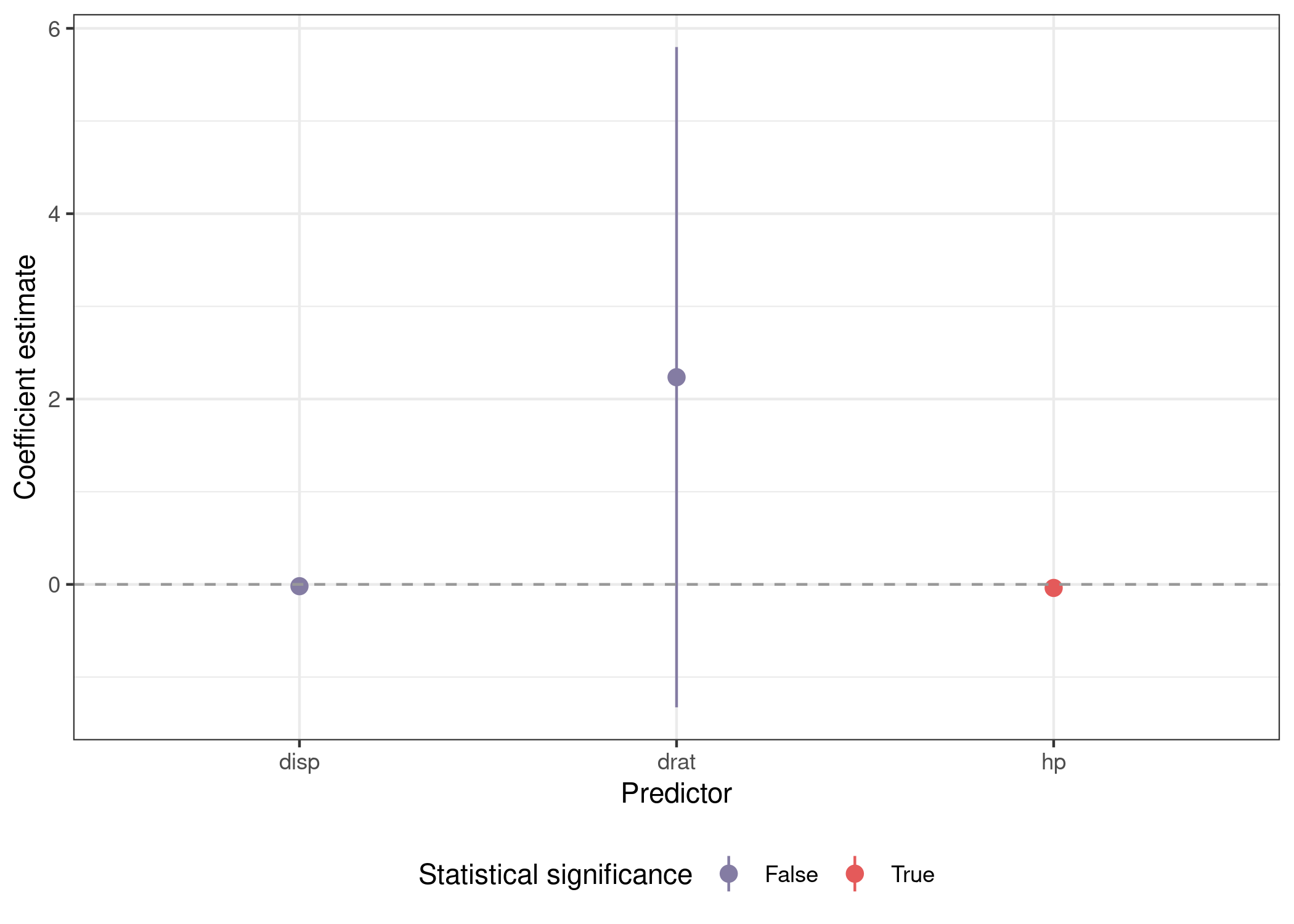
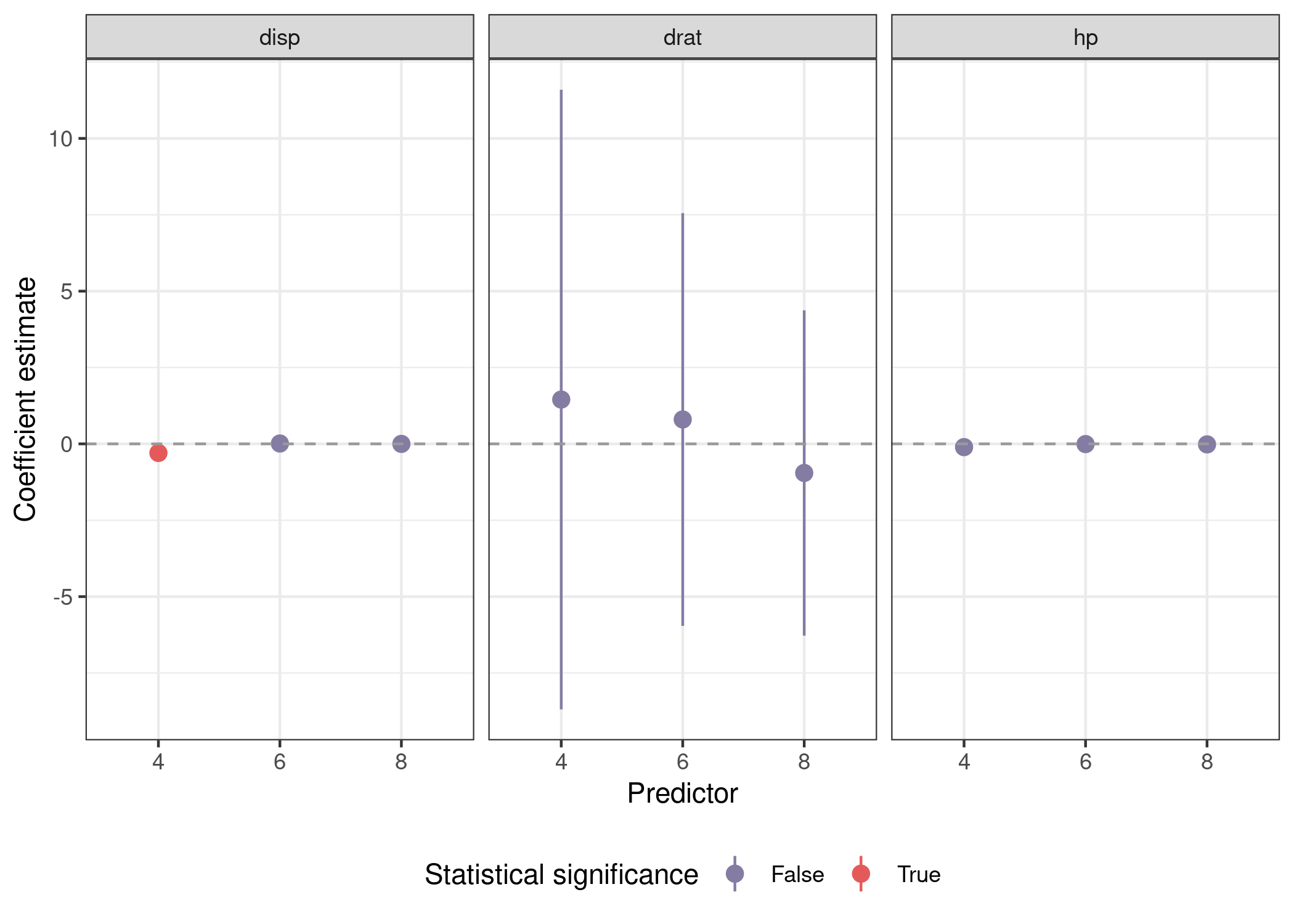
Plotting involves a call to facet_wrap(), but is otherwise similar.
mods_var_obs %>%
unnest(out) %>%
mutate(sig = sign(conf.low) == sign(conf.high)) %>%
filter(term %in% preds) %>%
ggplot(aes(x = cyl, y = estimate, ymin = conf.low, ymax = conf.high,
color = sig)) +
geom_pointrange() +
geom_hline(yintercept = 0, lty = 2, color = 'grey60') +
facet_wrap(~pred) +
scale_color_manual(name = 'Statistical significance',
labels = str_to_title,
values = park_palette('Saguaro')) +
labs(x = 'Predictor', y = "Coefficient estimate") +
theme_bw() +
theme(legend.position = 'bottom')
Creating tables is more complex. Here we have to cycle through each
predictor with a call to map(), filter the output to only contain
results from models using that predictor, then split the dataframe by
cylinders instead of into separate rows. Note the use of
unname(preds_name[x]) to retrieve full english predictor names to
create more useful table titles. We’ll also be using tab_coef_map from
above to get more informative row labels in our tables. Running the code
below generates the following tables:
## named vector for full english predictor names
preds_name <- c('displacement', 'horsepower', 'drive ratio')
names(preds_name) <- preds
map(preds, function(x) mods_var_obs %>%
filter(pred == x) %>% # subset to models using predictor x
select(mod, cyl) %>% # drop tidied model
split(.$cyl) %>% # split by number of cylinders in engine
lapply(function(y) y$mod[[1]]) %>% # only one item in each list
modelsummary(title = str_c('Predictor: ',
unname(preds_name[x]), # formatted name
coef_map = tab_coef_map,
stars = c('*' = .05),
escape = F) %>%
add_header_above(c(' ' = 1, 'Cylinders' = 3))) %>%
walk(print) # invisibly return input to avoid [[1]] in output
We’ve got one table for each predictor we considered, and each one is
split into three models for cars with four, six, and eight cylinder
engines. This is a bit overkill for this example, but it’s all you have
to do to scale this framework up to hundreds of potential predictors is
put more items in preds.
-
Yes, I know this is a perfect situation to use LASSO. Sometimes people (reviewers) want certain models run, and you just have to run them. ↩
-
There’s a very real chance that someone else is me in six months. ↩
-
Things get a lot more complicated if your
split()call produces a list of dataframes that aren’t one row each, so make sure that’s what you’re getting before you proceed. ↩

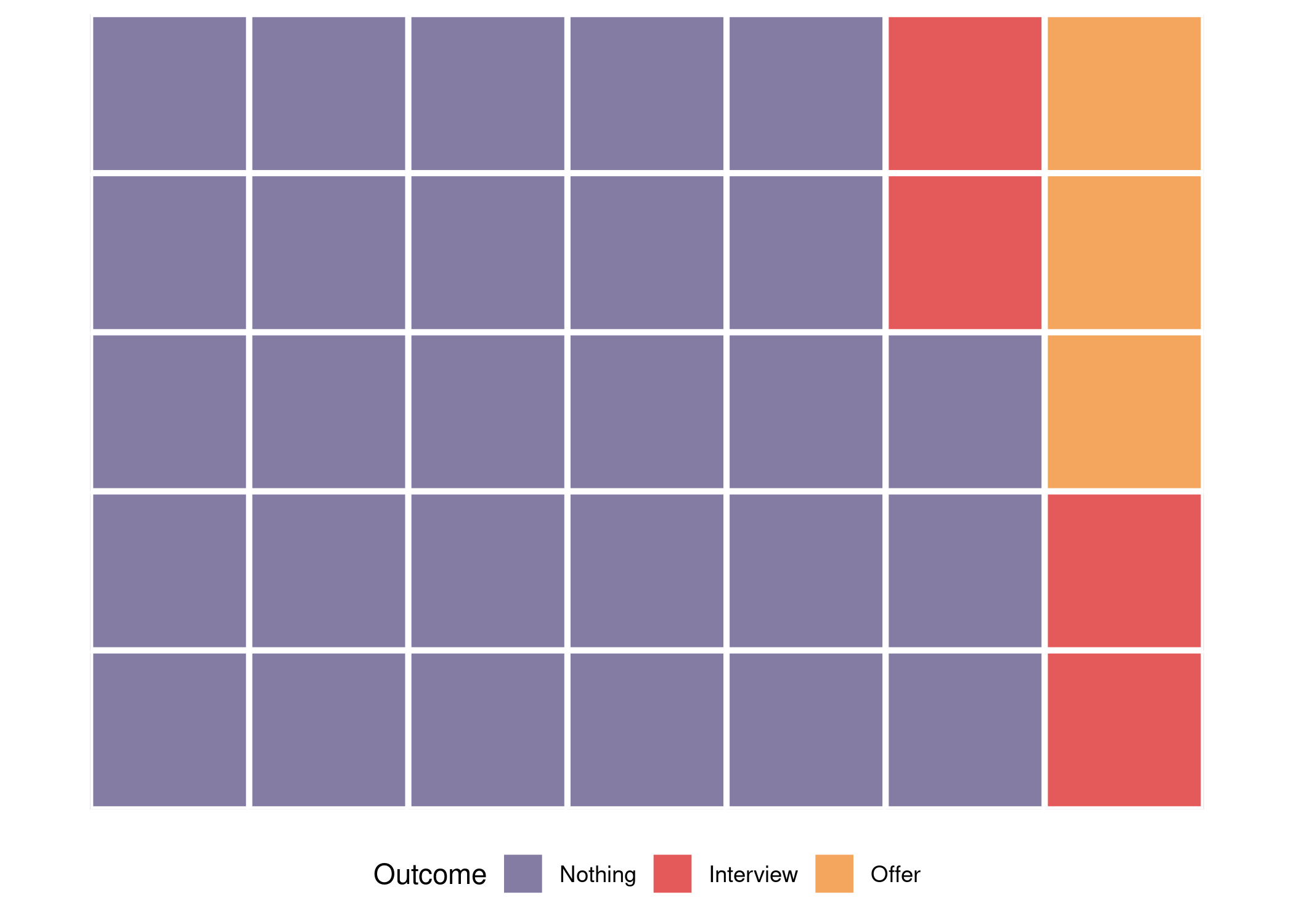
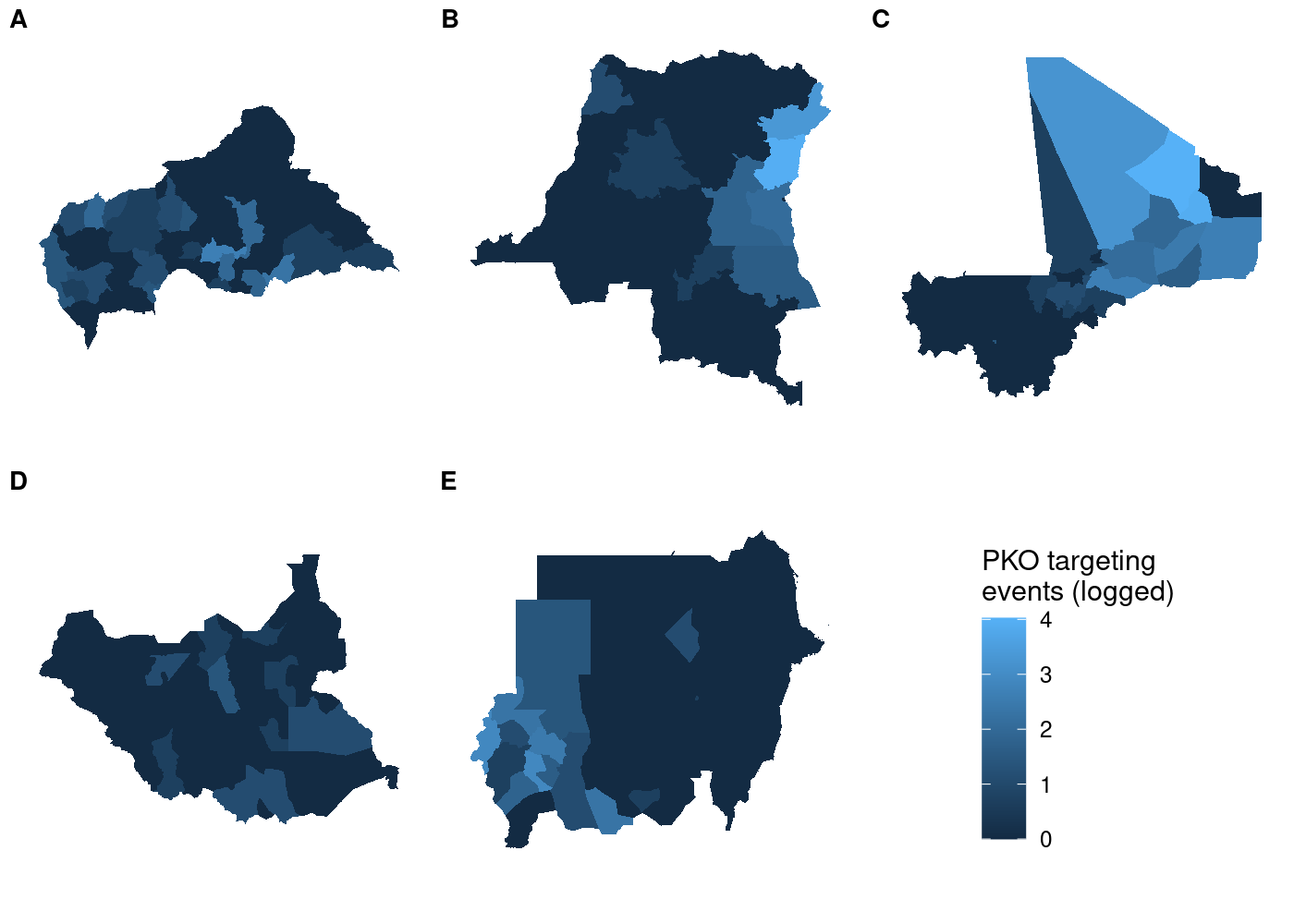
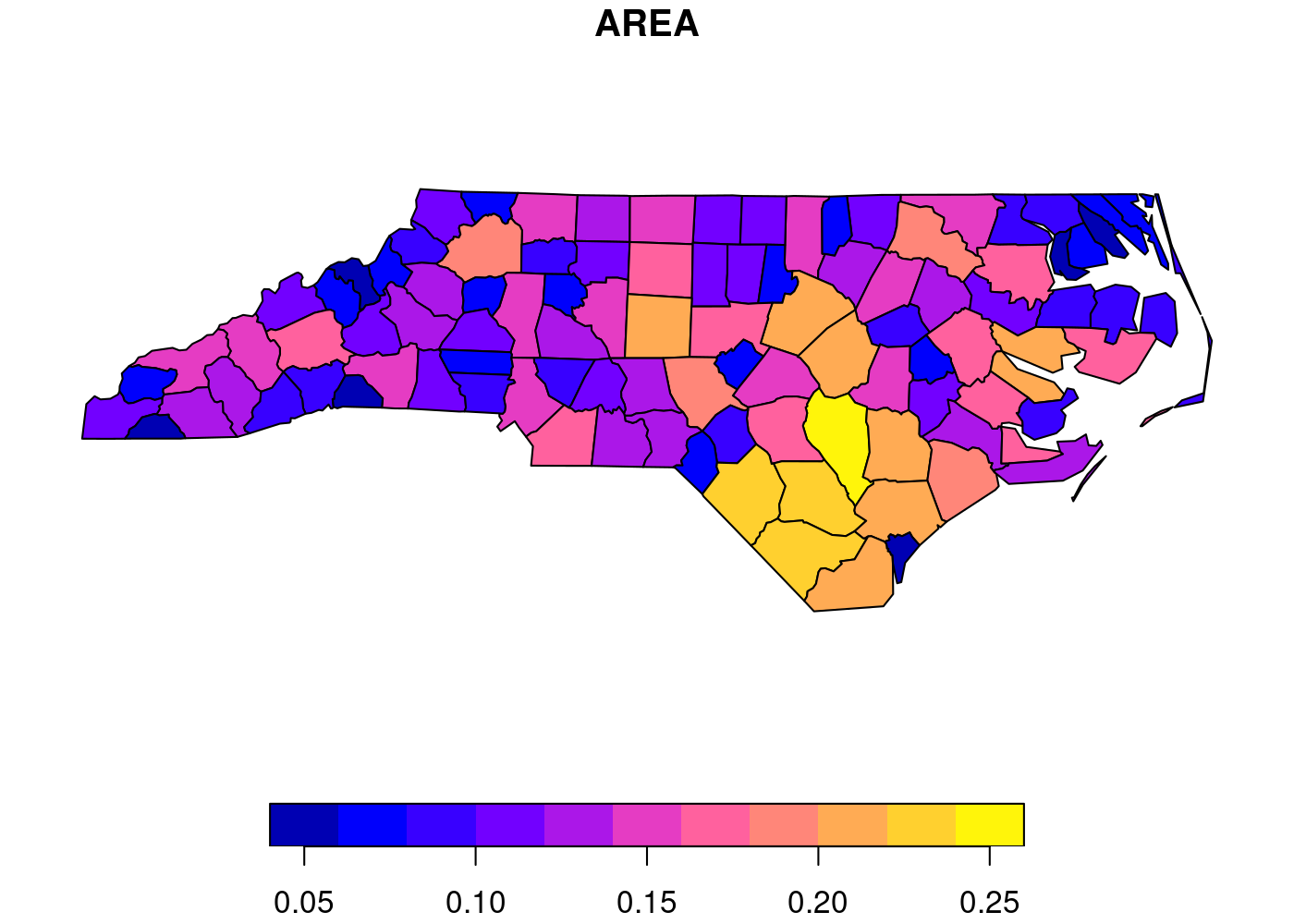 Since I needed to generate choropleths for multiple countries, I decided
to use
Since I needed to generate choropleths for multiple countries, I decided
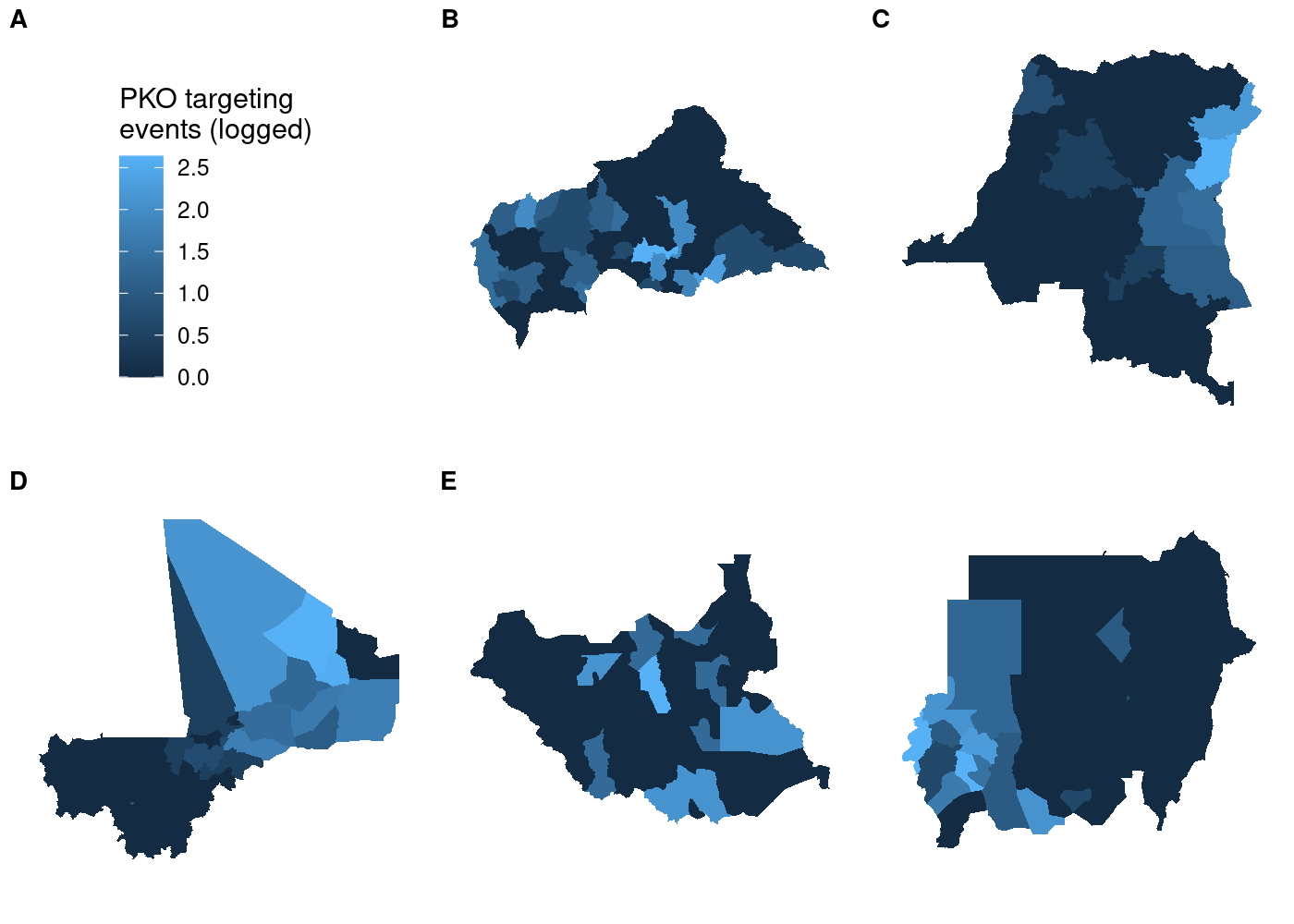
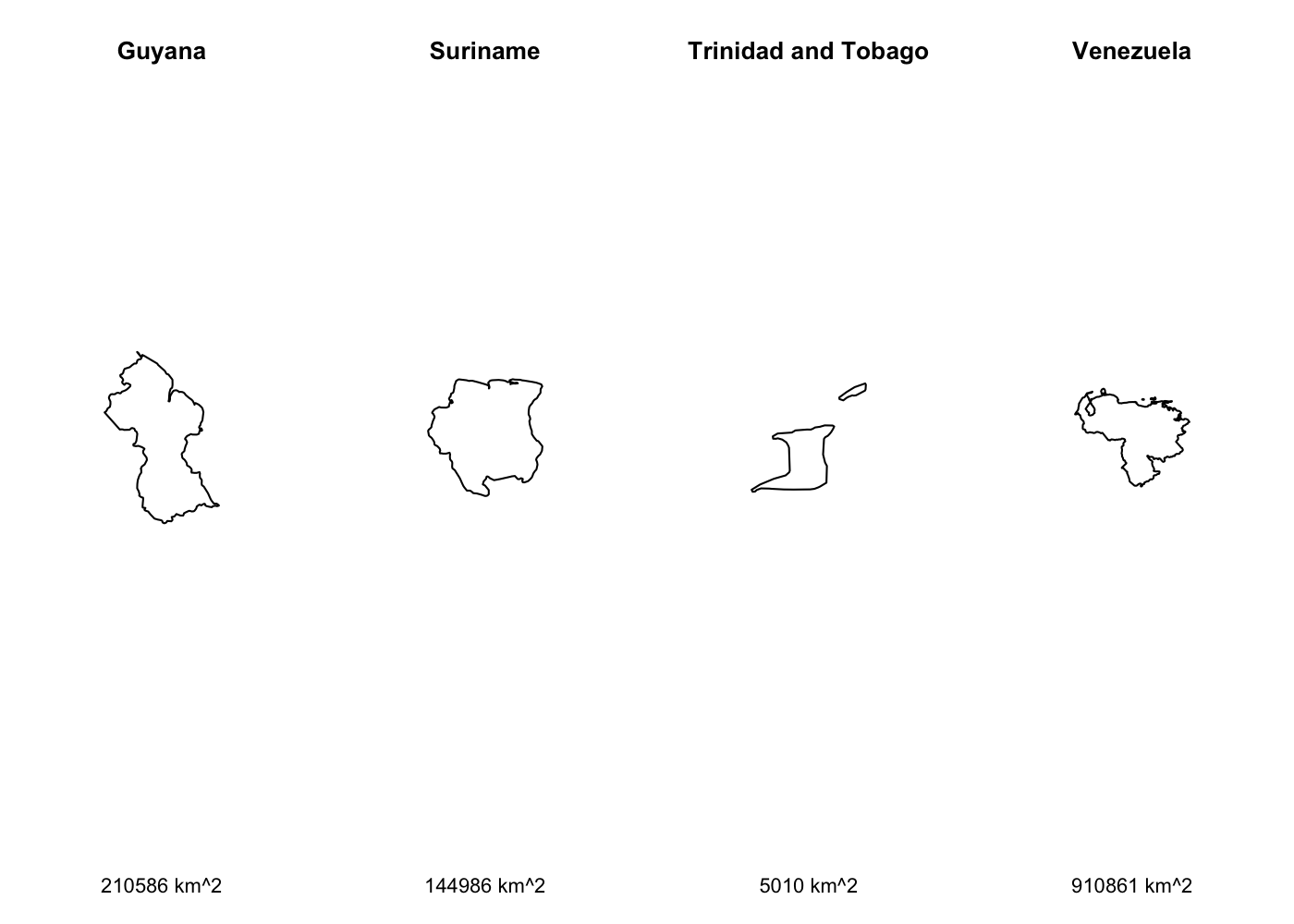
to use 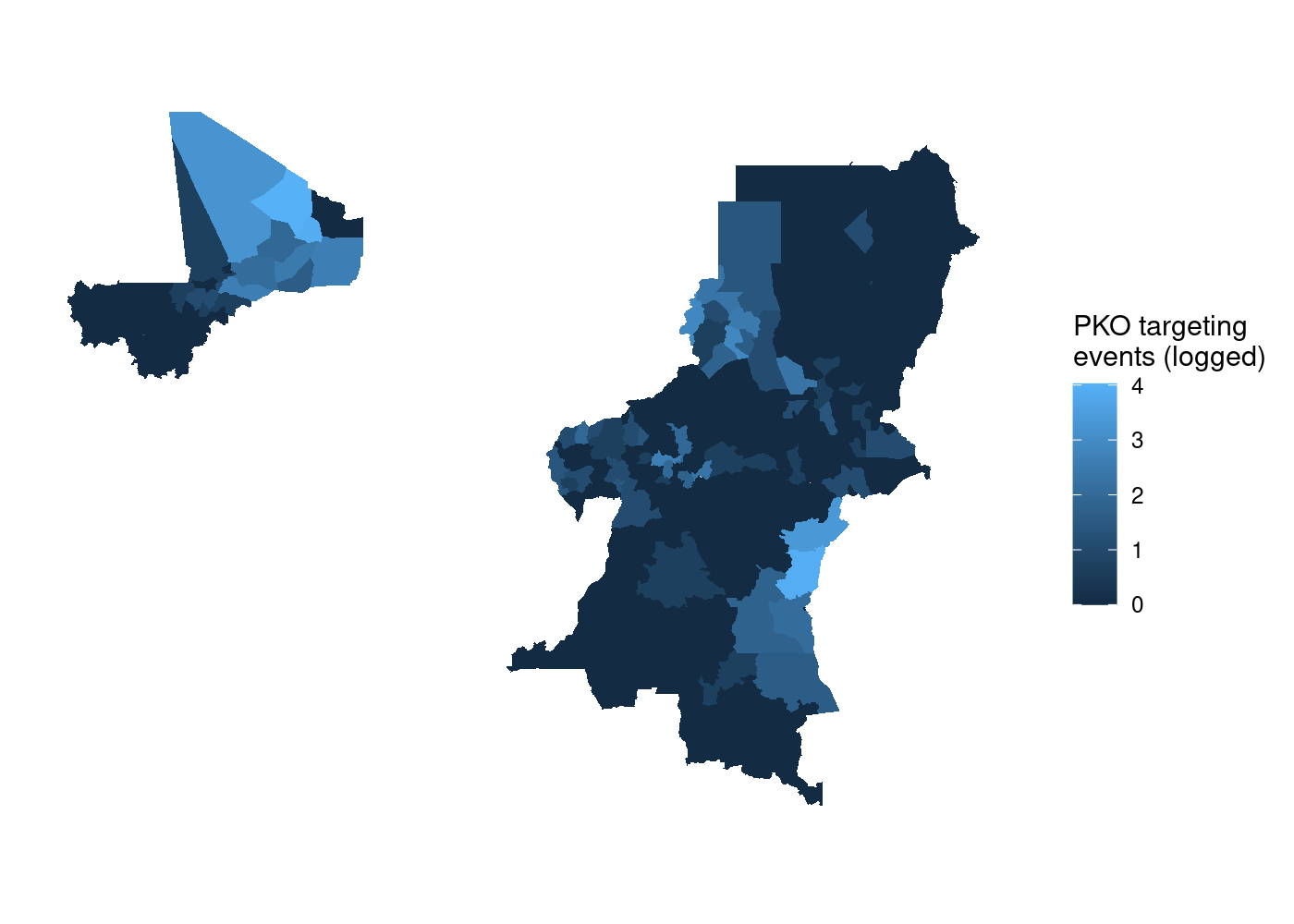 That’s a lot of wasted white space, and it can make certain countries
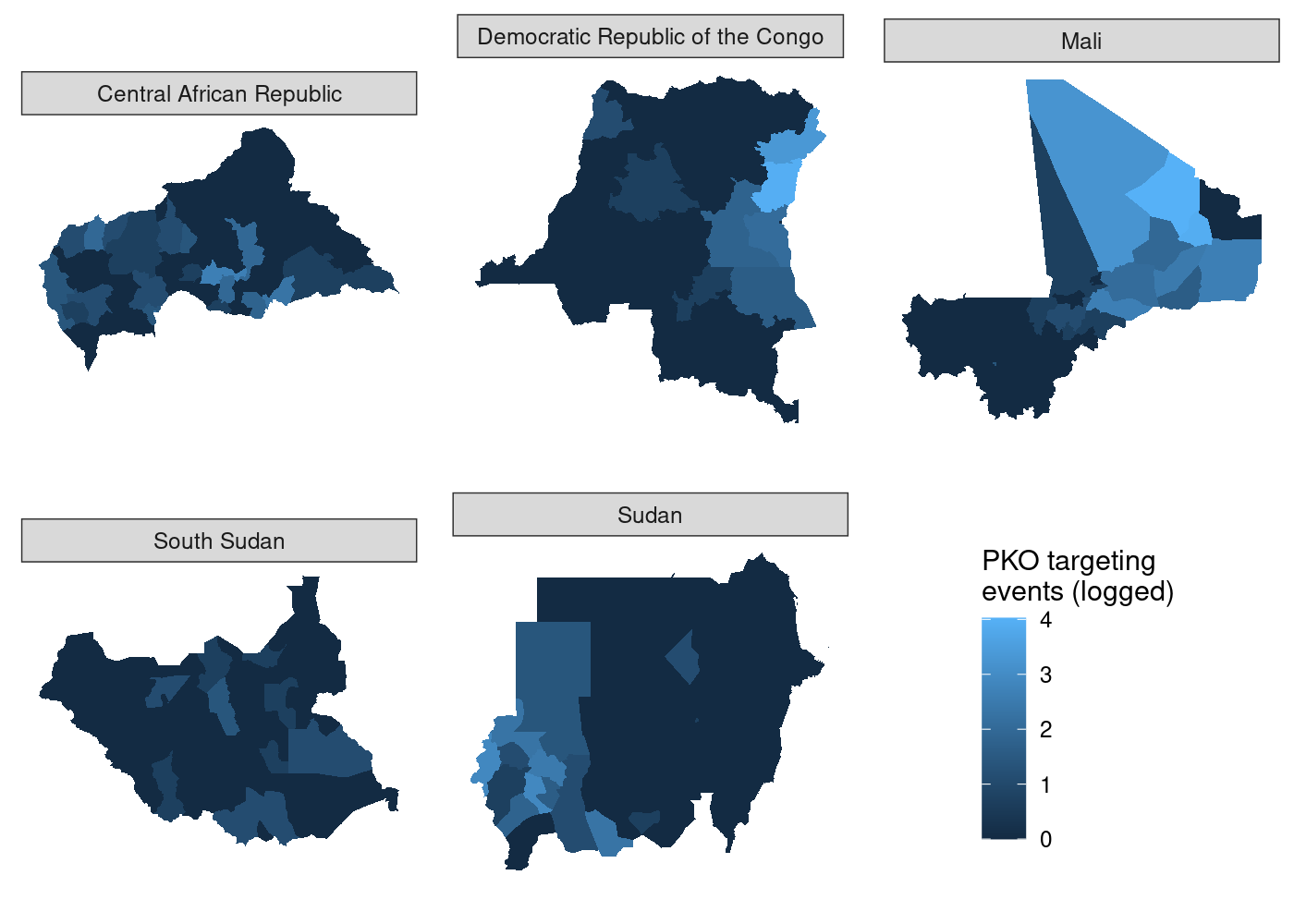
harder to see. Let’s split it out using
That’s a lot of wasted white space, and it can make certain countries
harder to see. Let’s split it out using 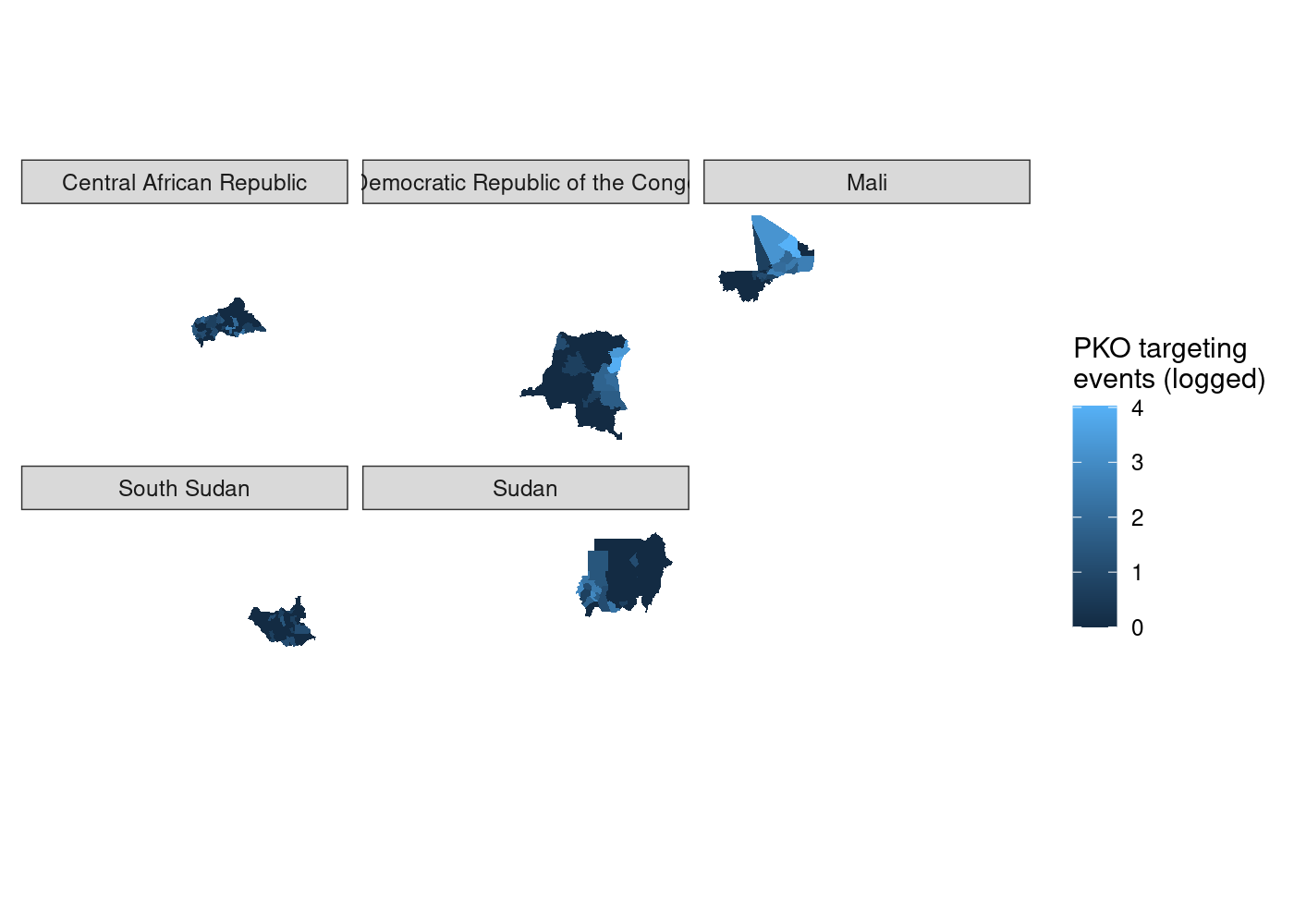 We’ve got facets, but everything is still clearly on the same scale.
let’s set
We’ve got facets, but everything is still clearly on the same scale.
let’s set 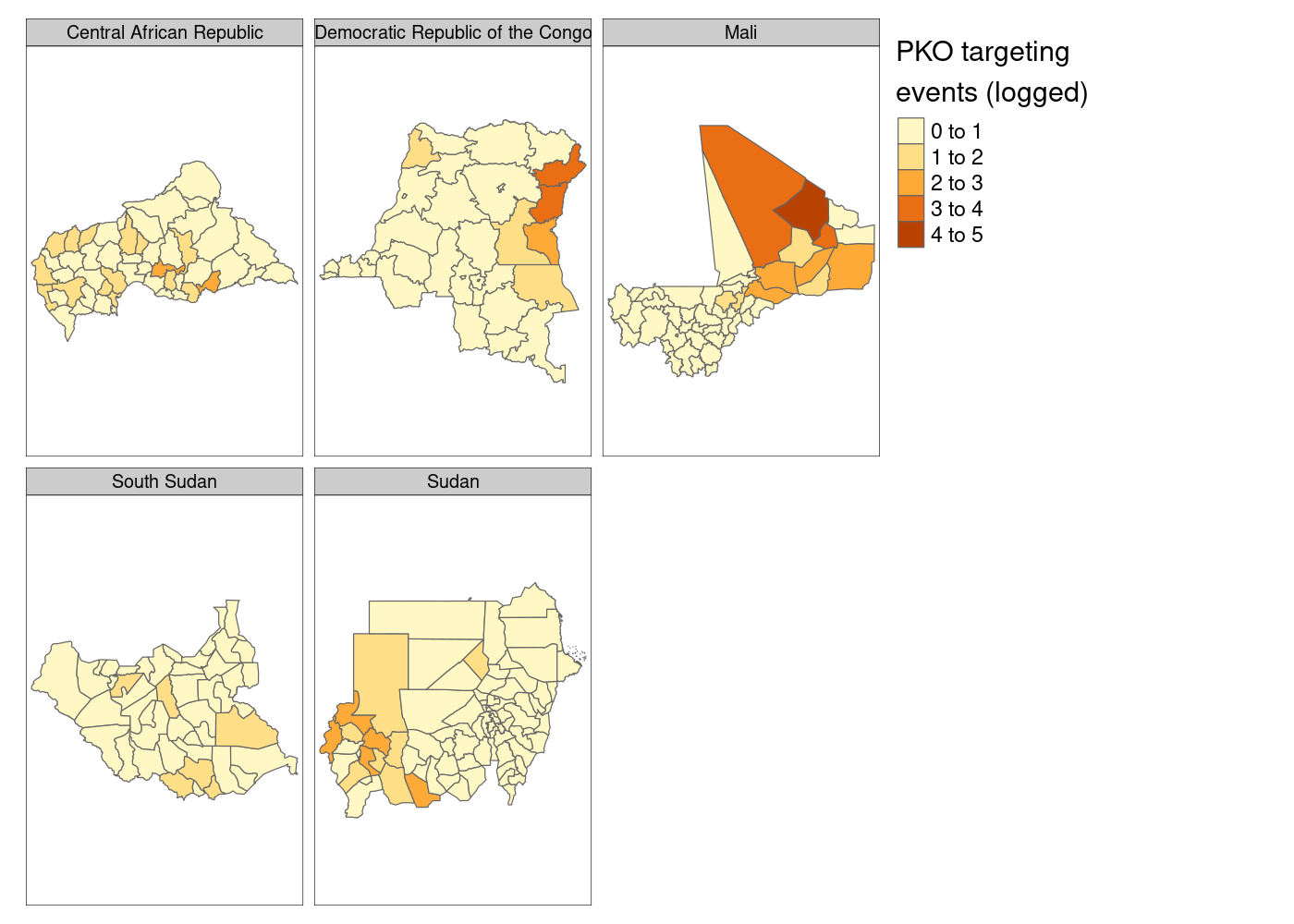
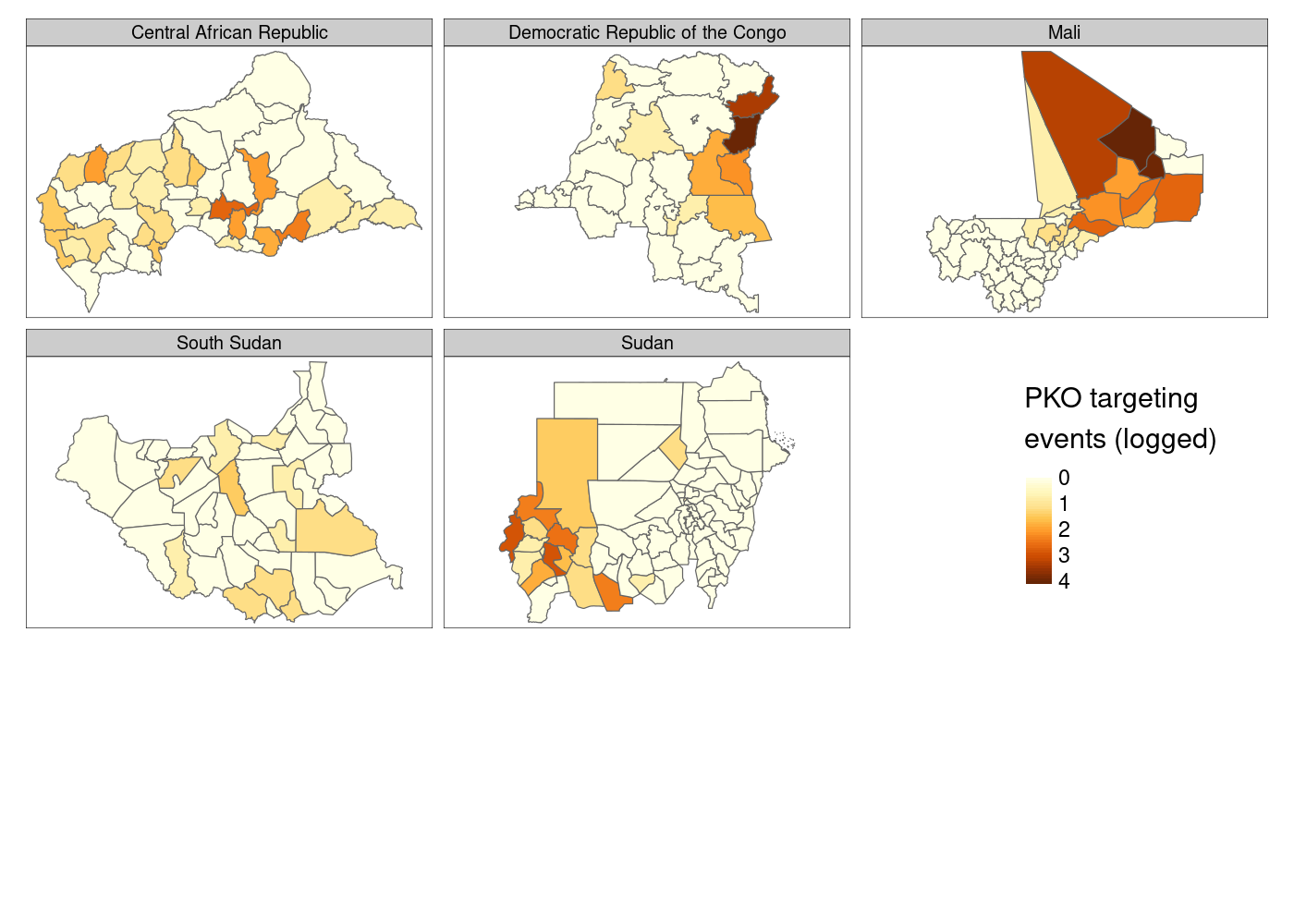
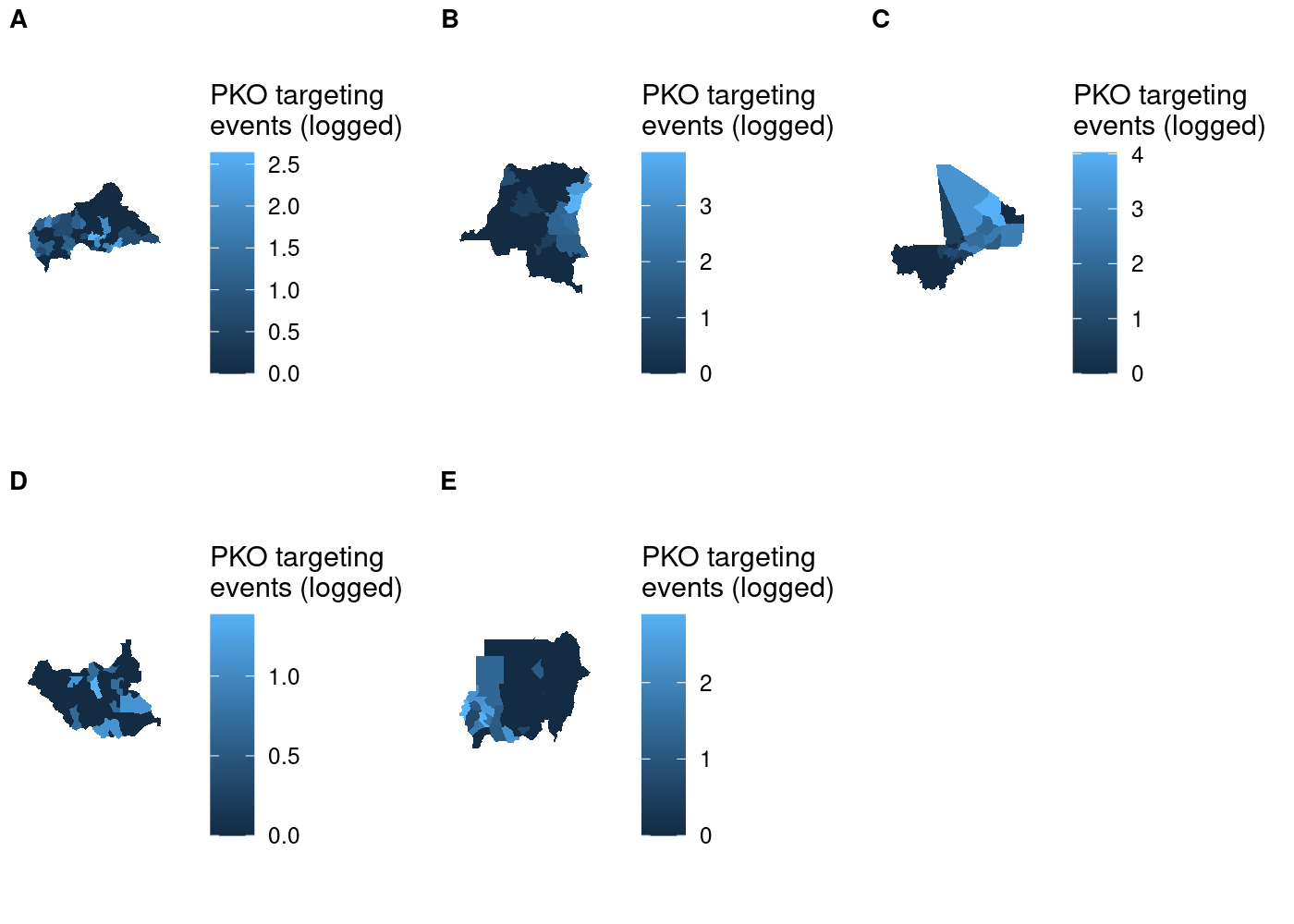
 This doesn’t look right! We told
This doesn’t look right! We told 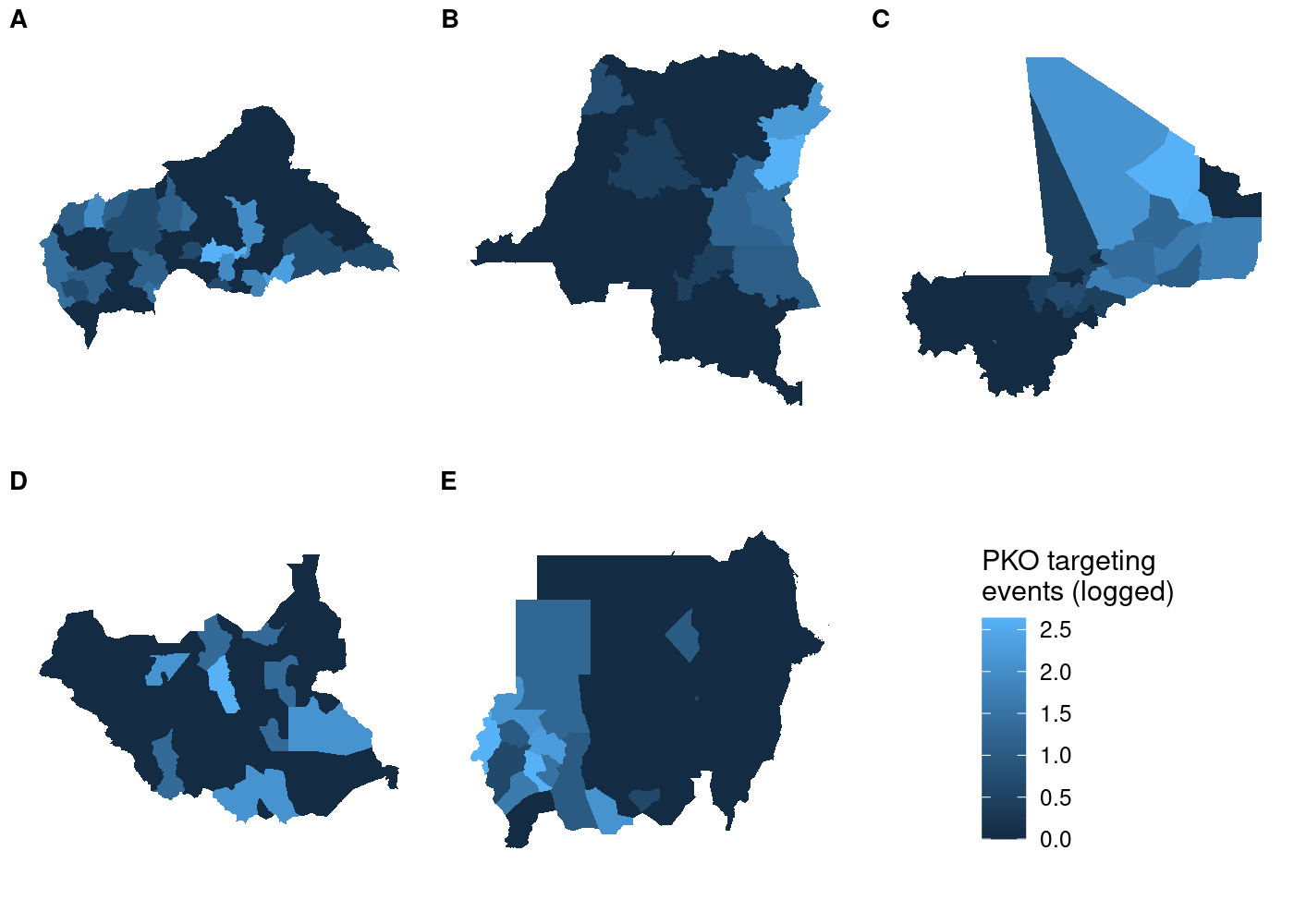
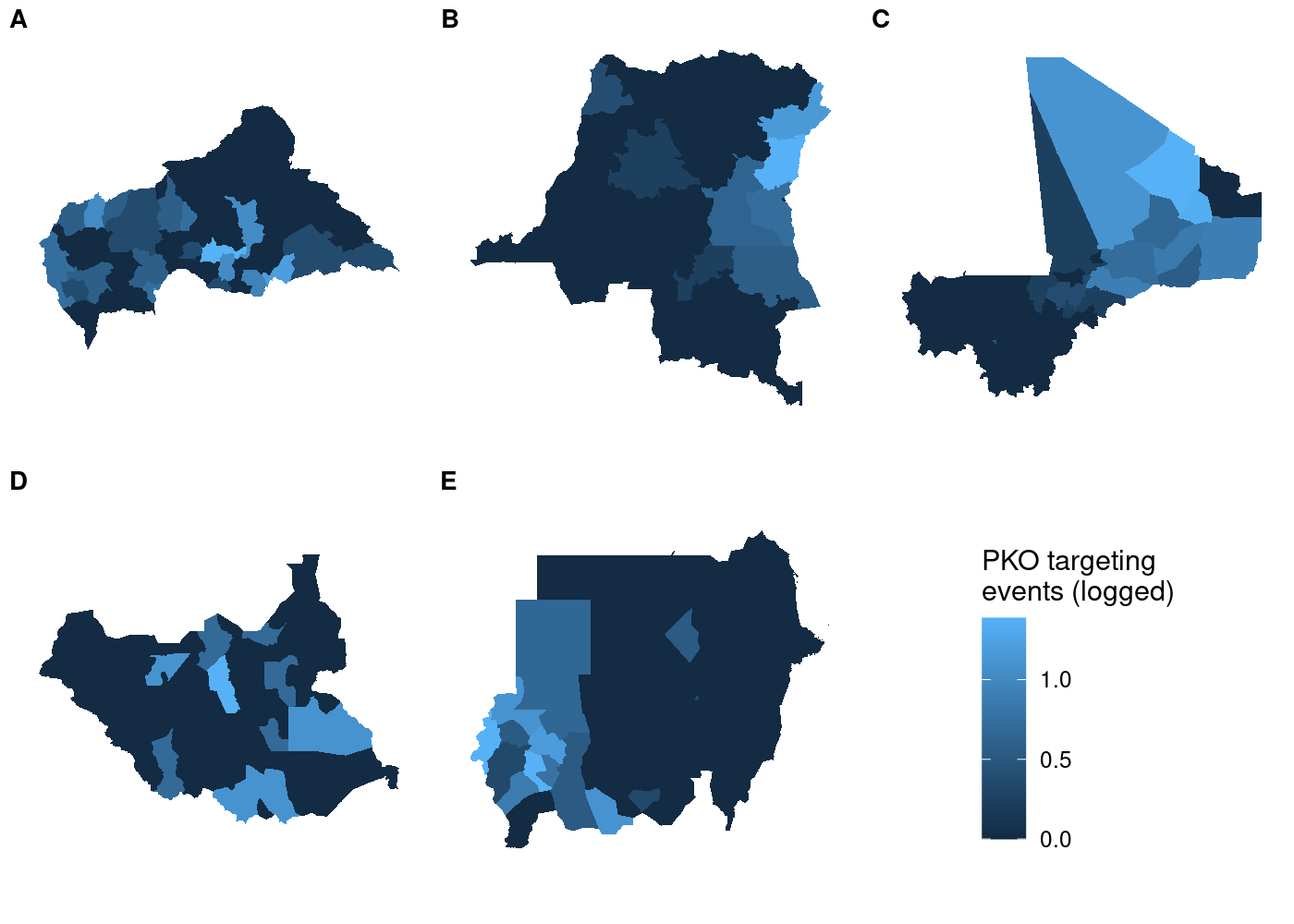 Each subplot has its own unique legend that’s automatically generated
from the values of
Each subplot has its own unique legend that’s automatically generated
from the values of 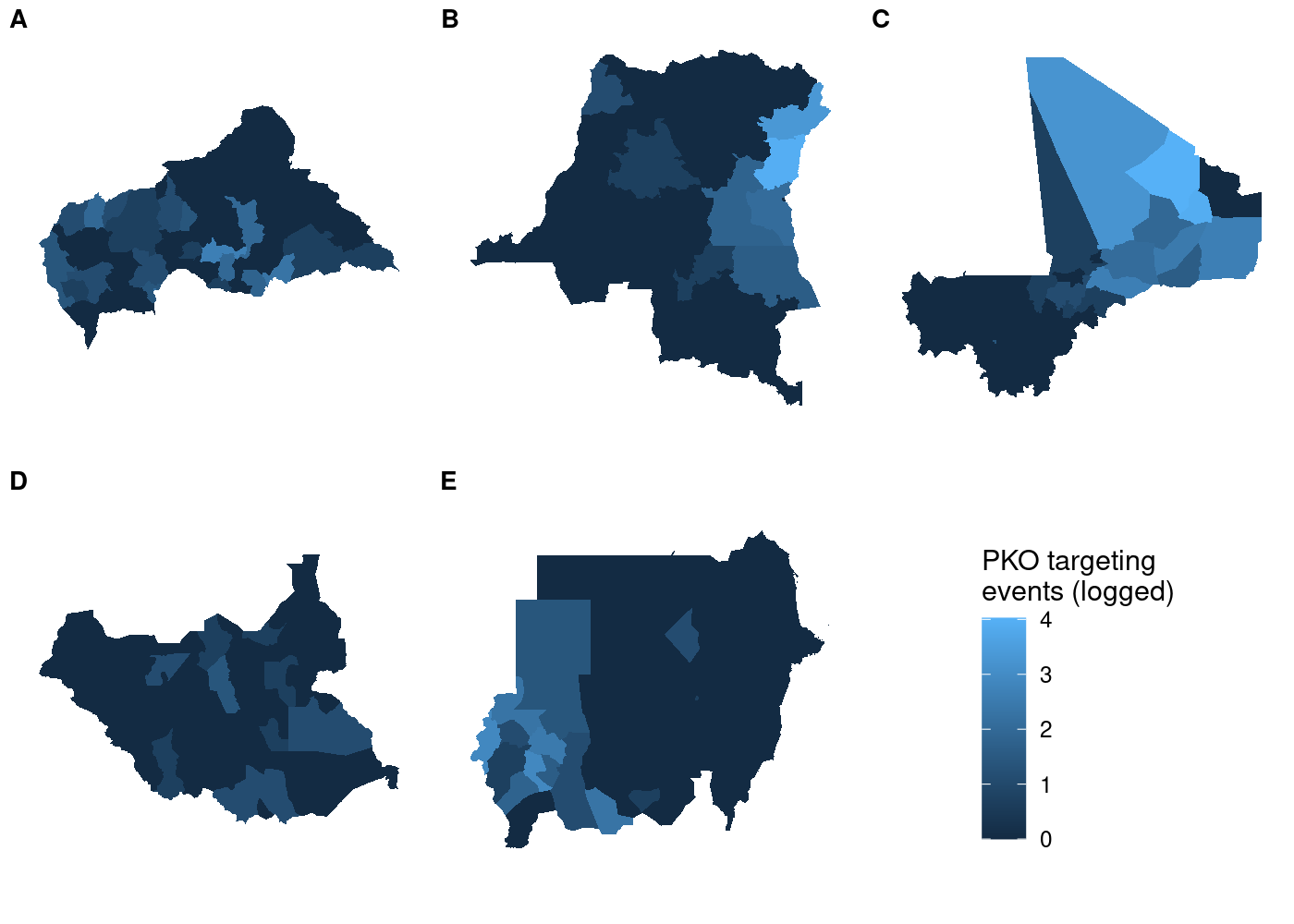
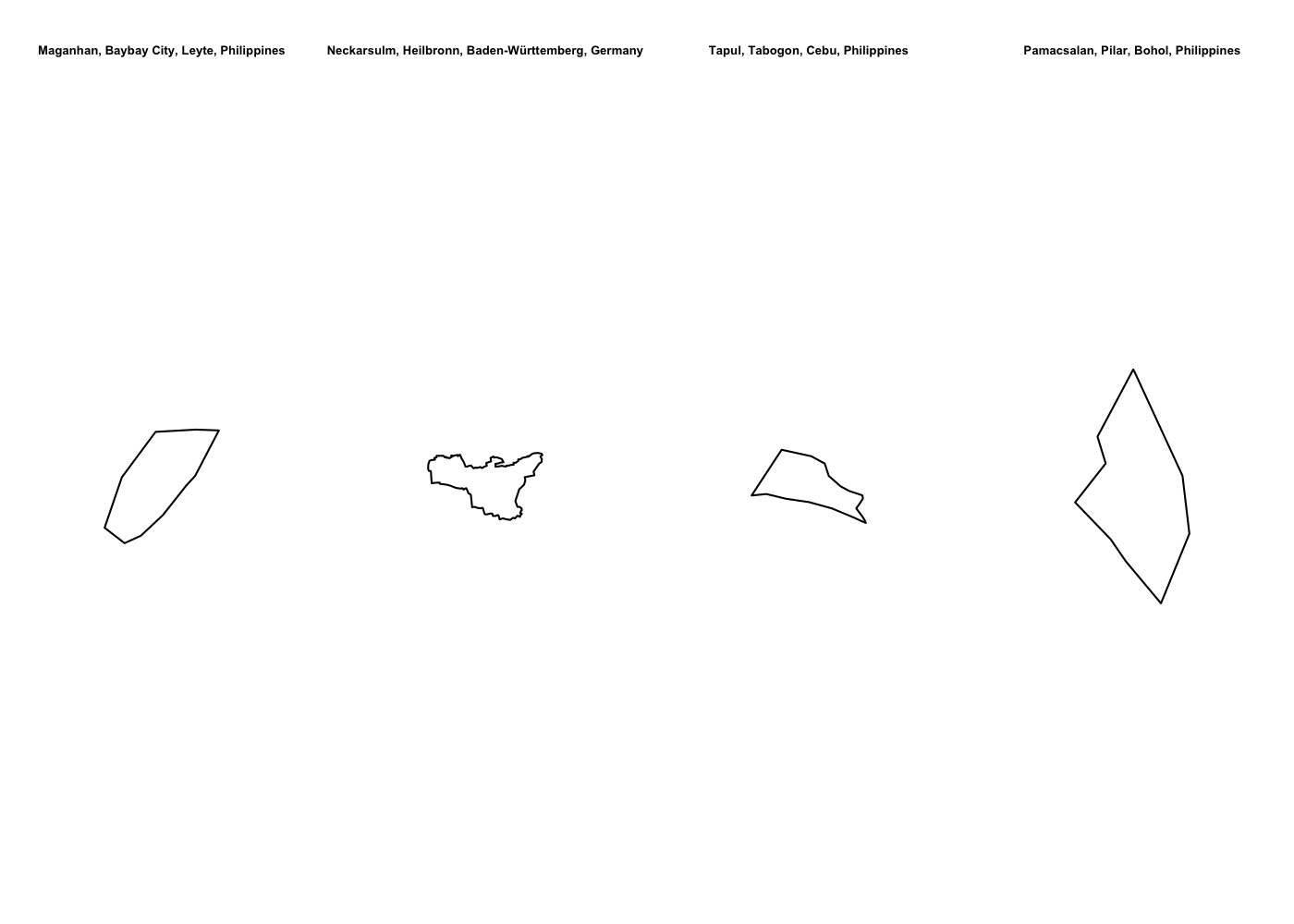
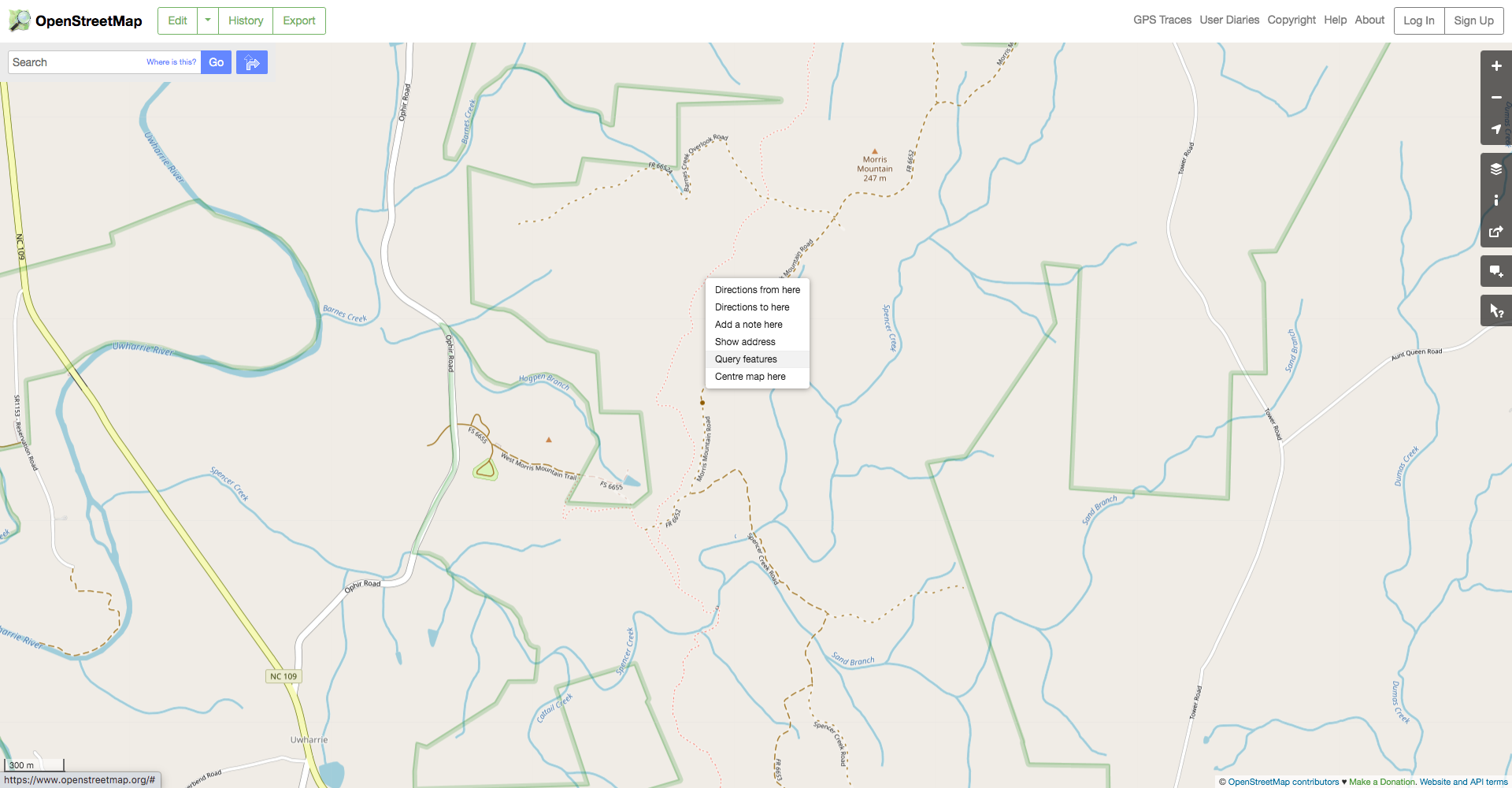
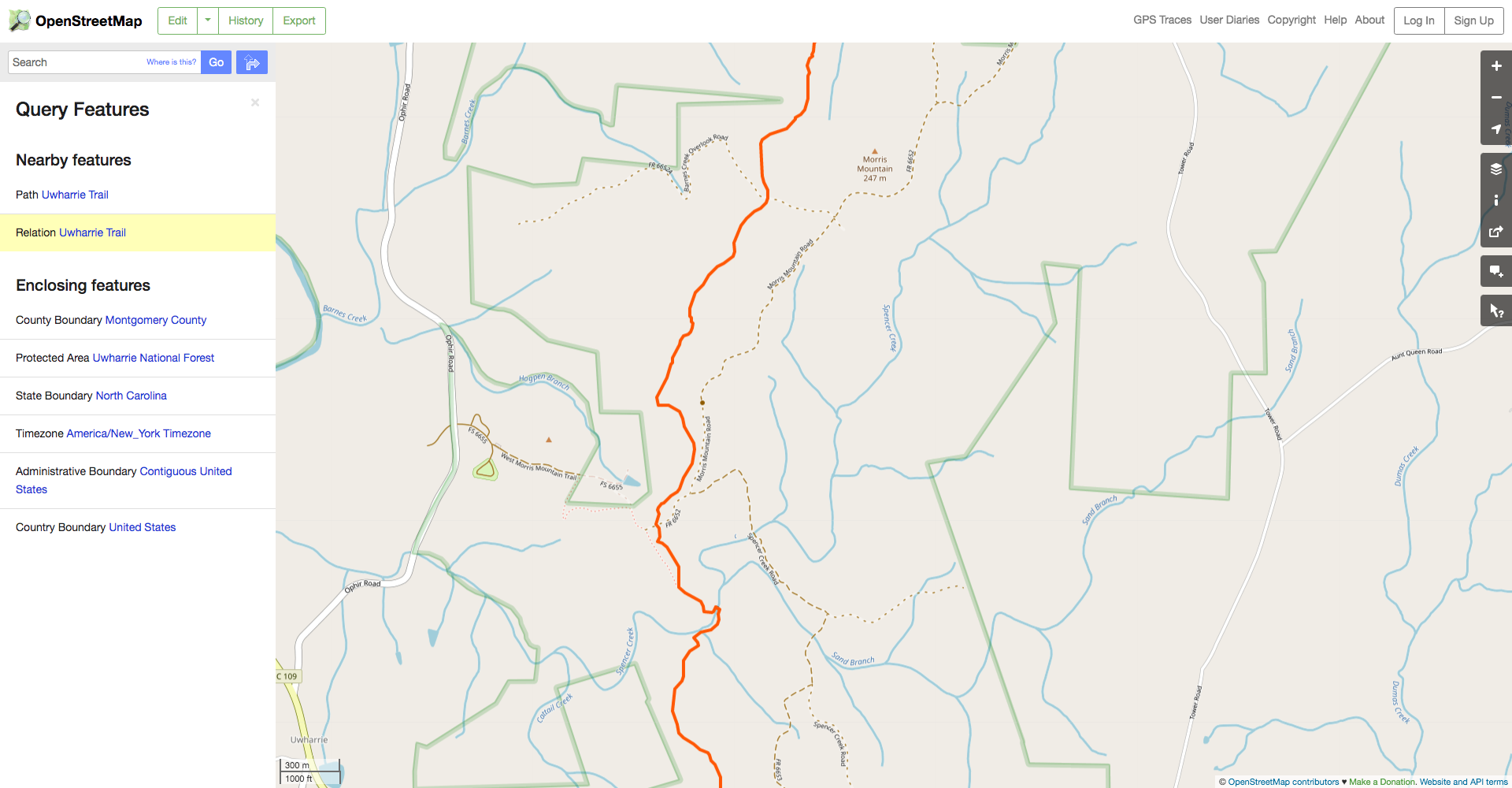
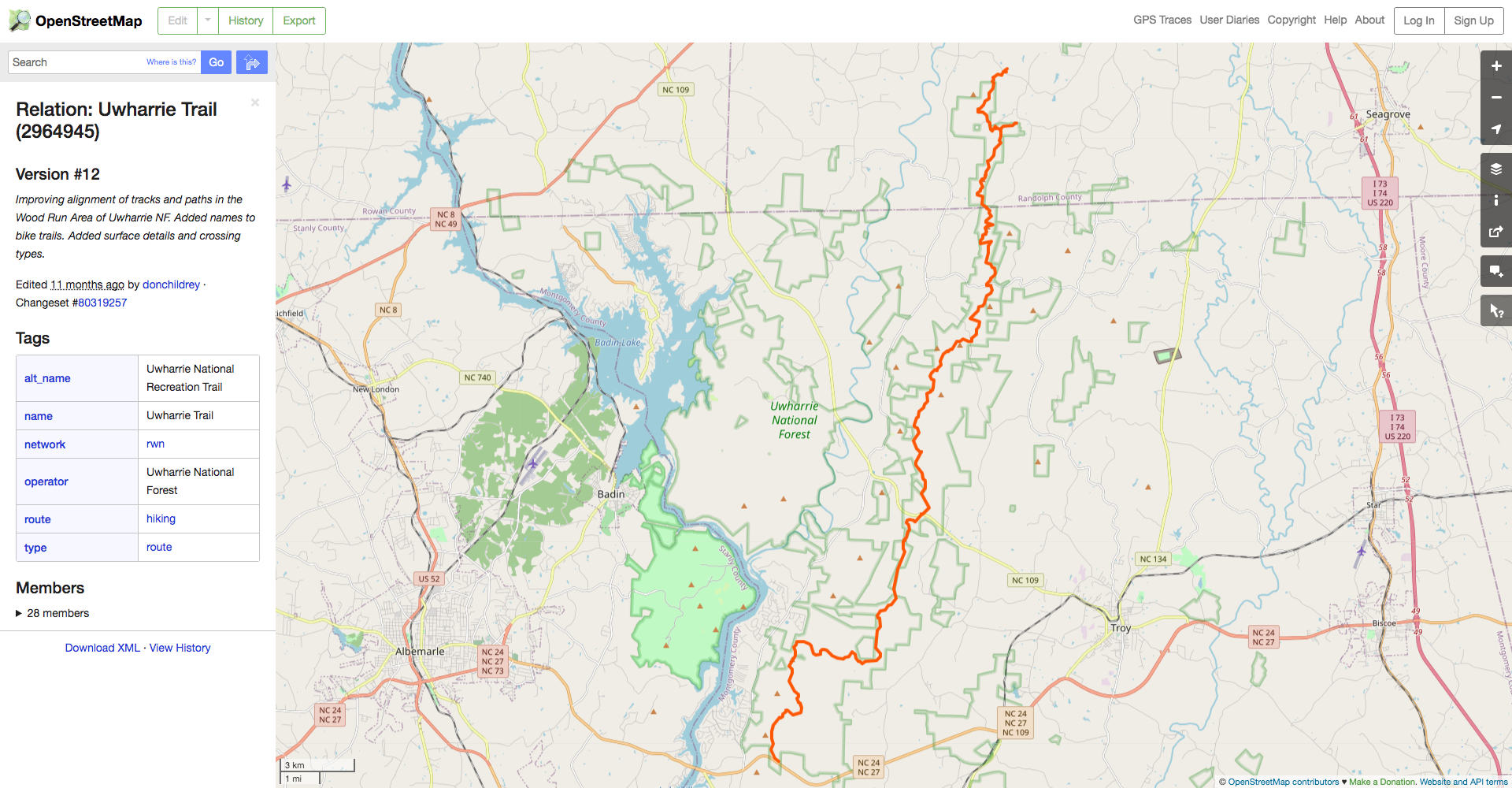
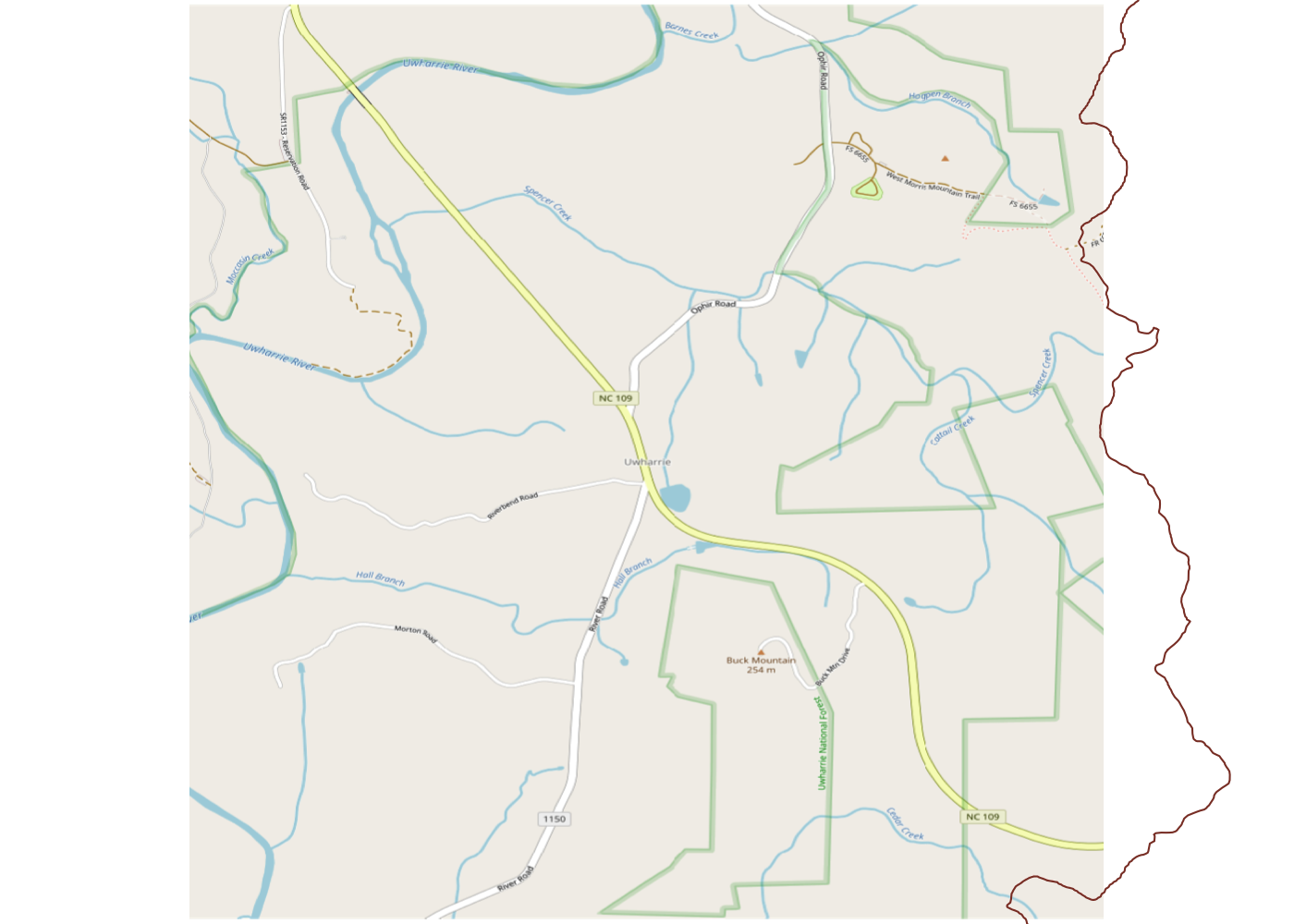
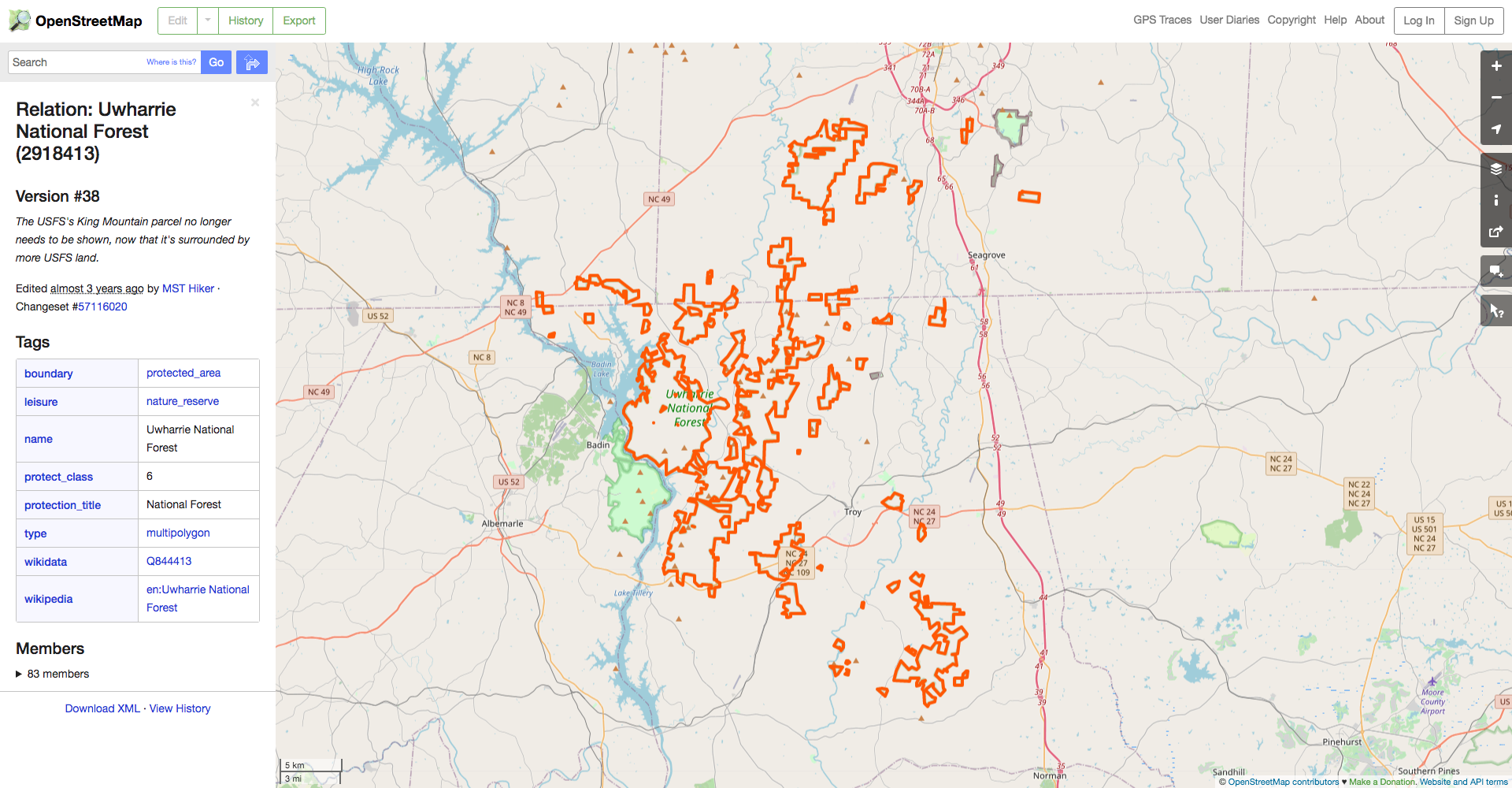
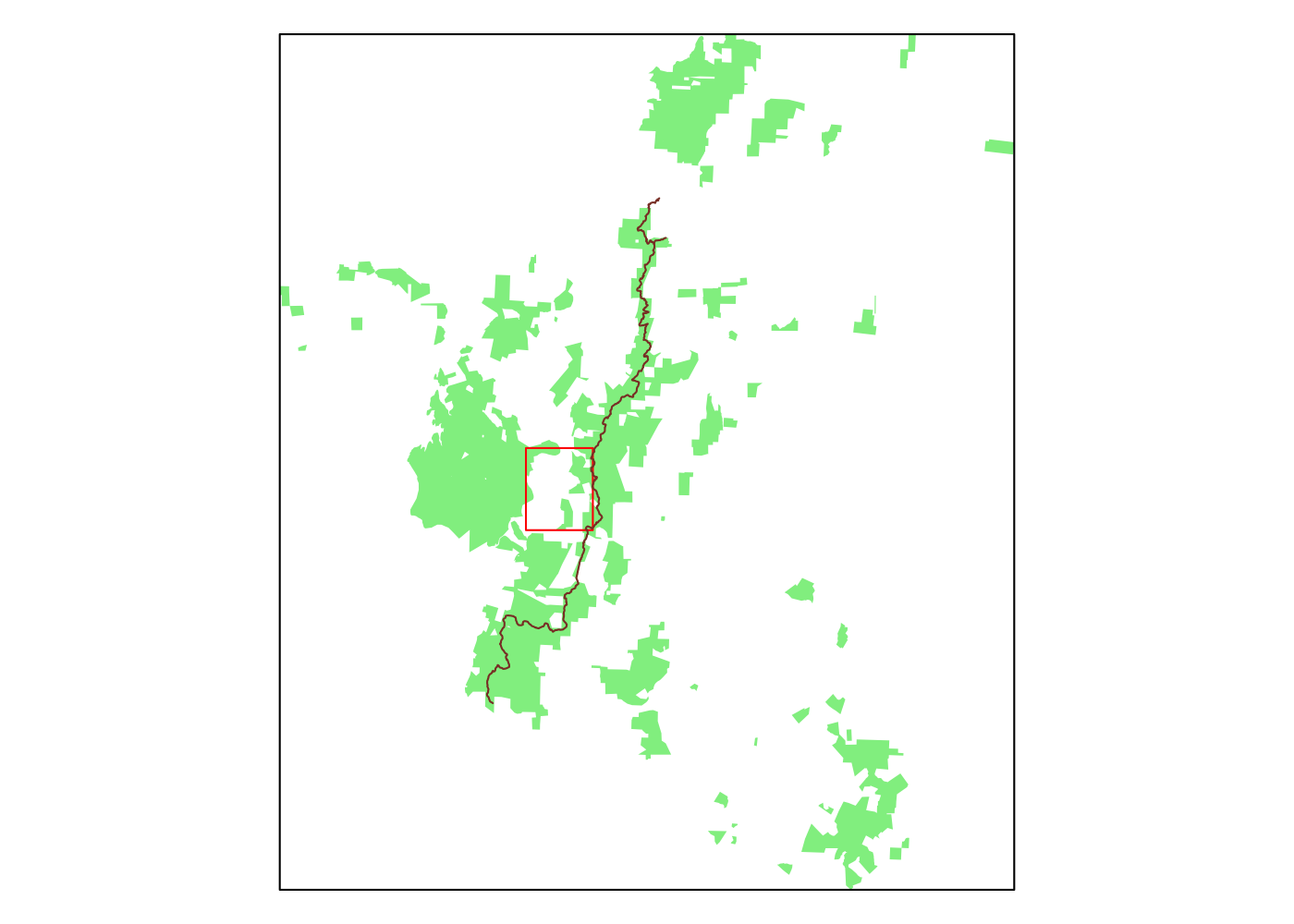
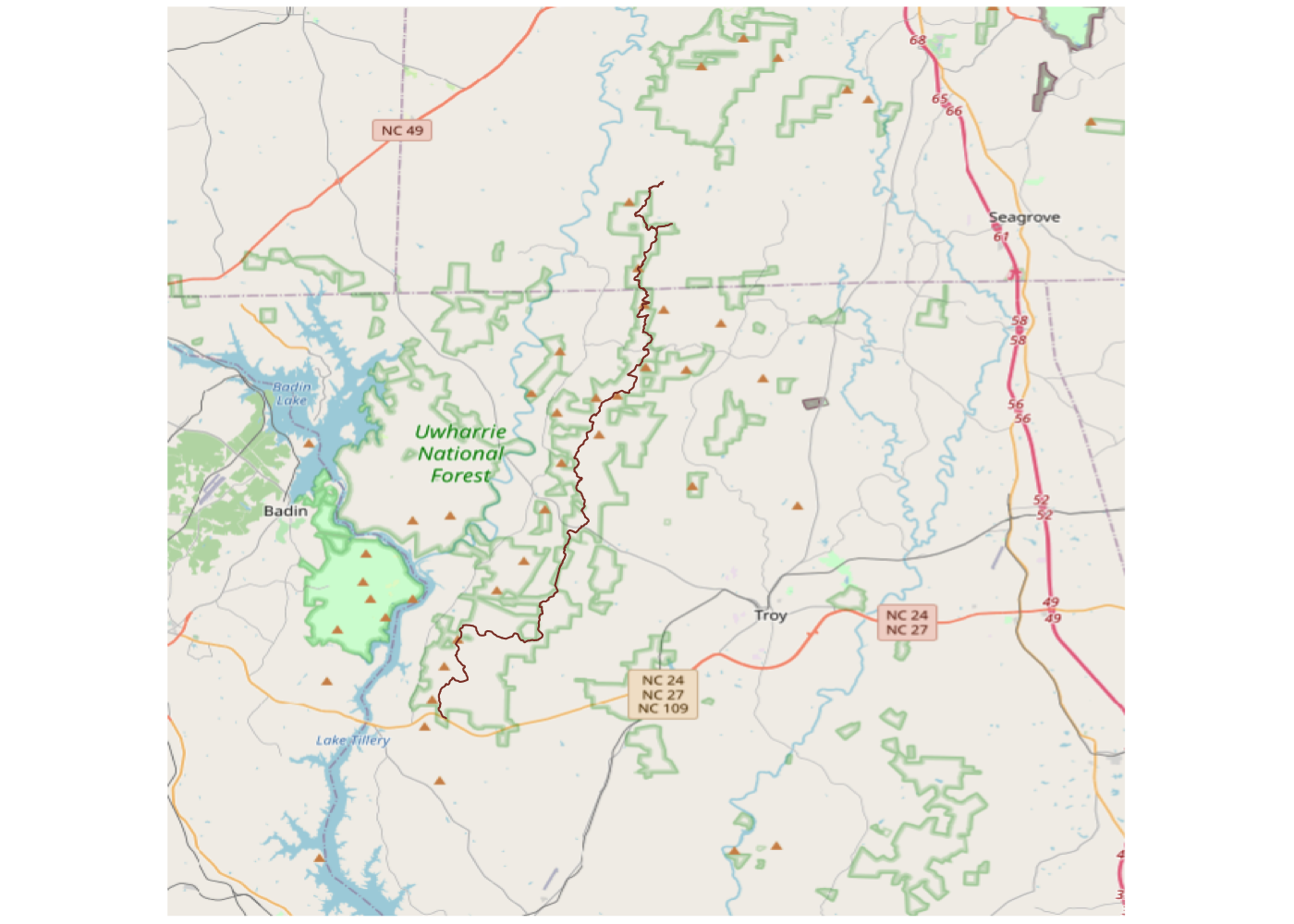
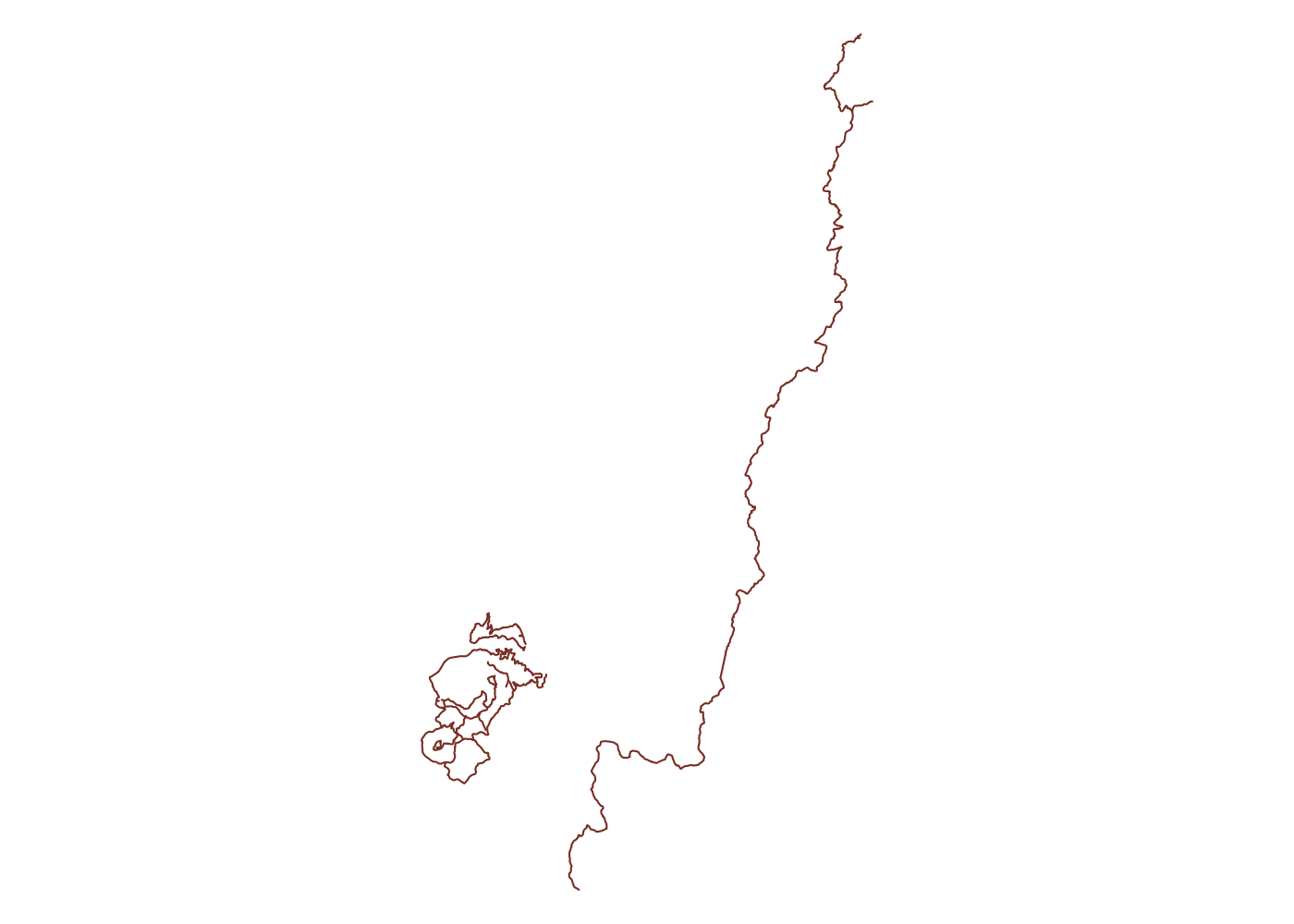
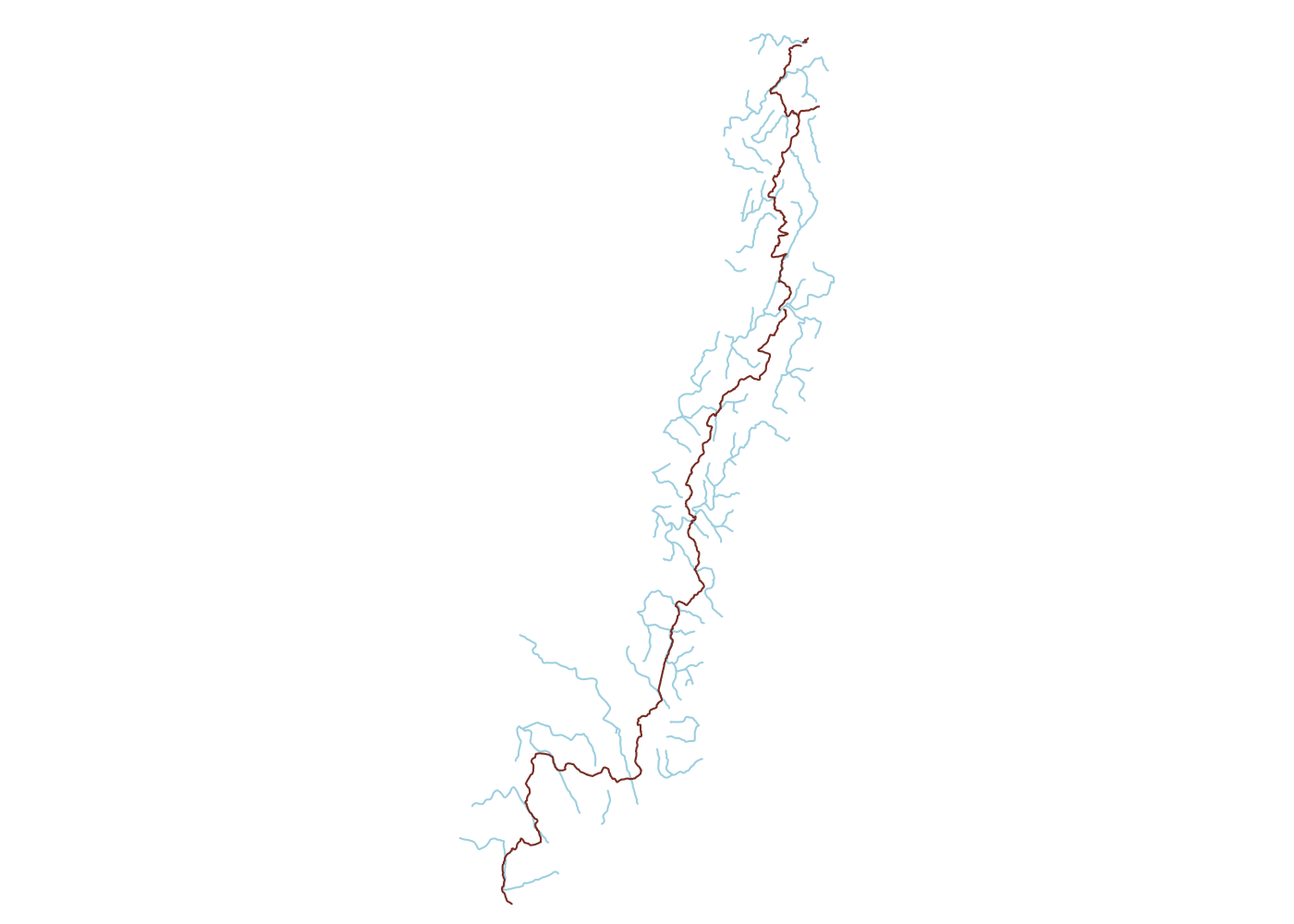
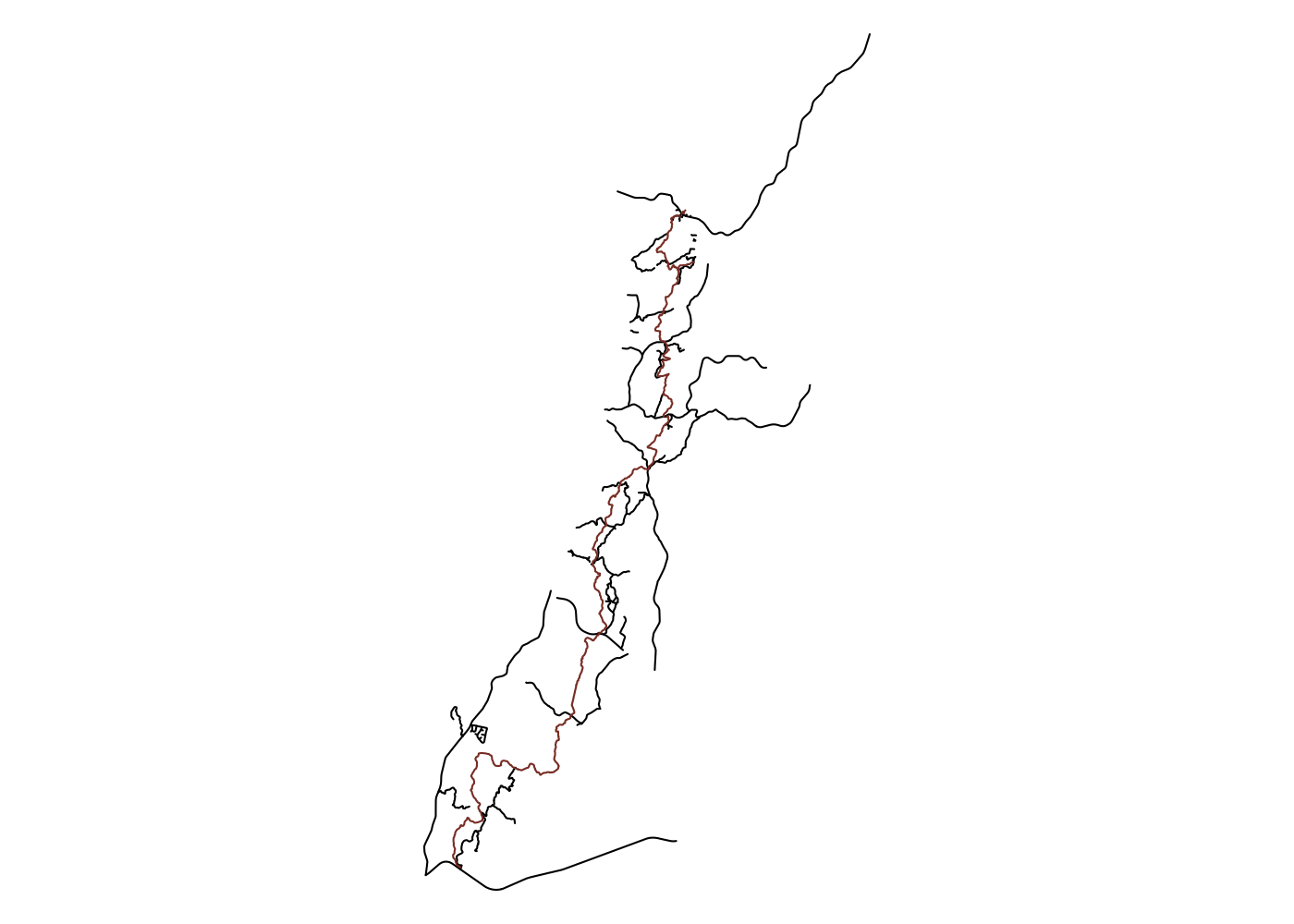

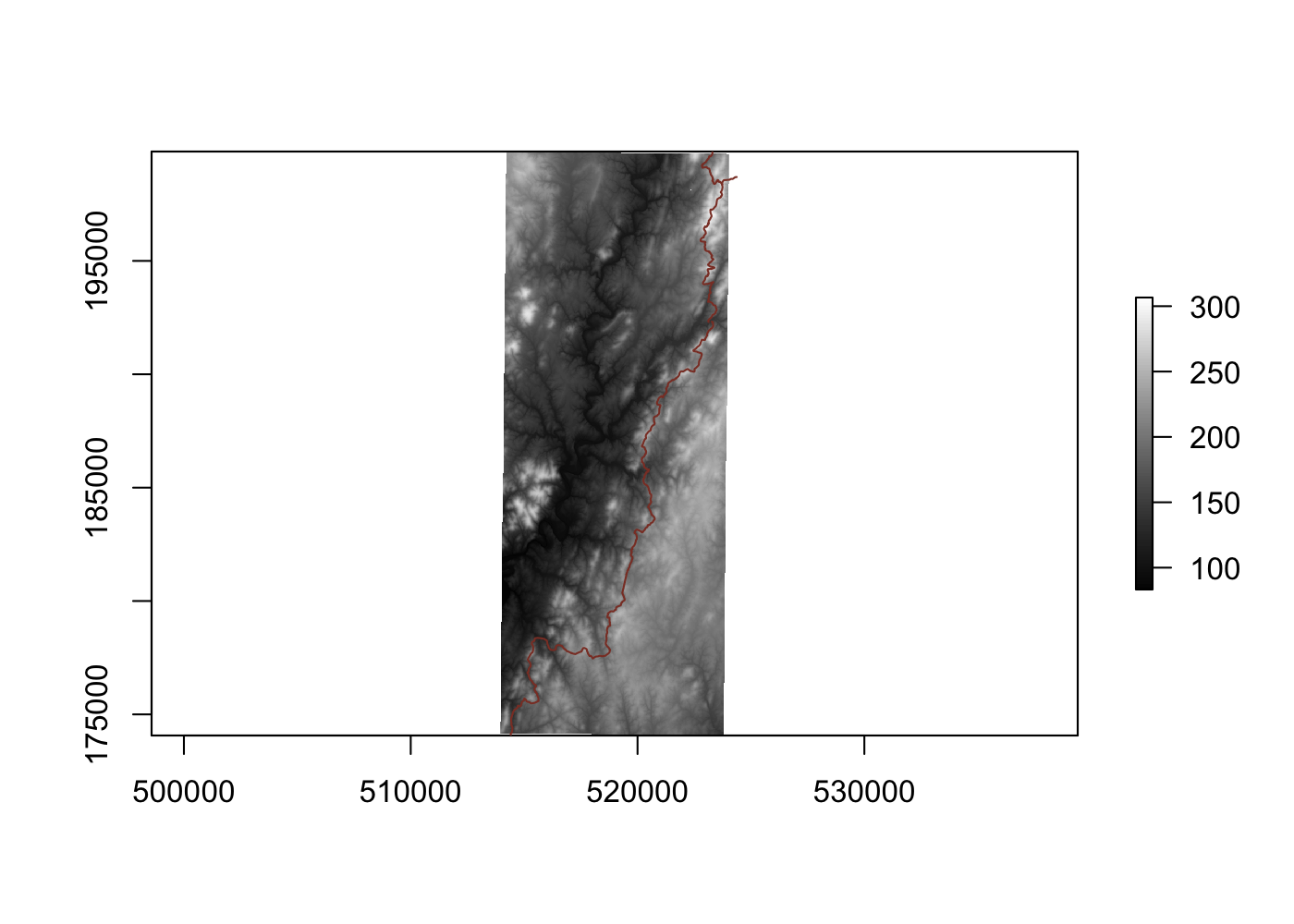
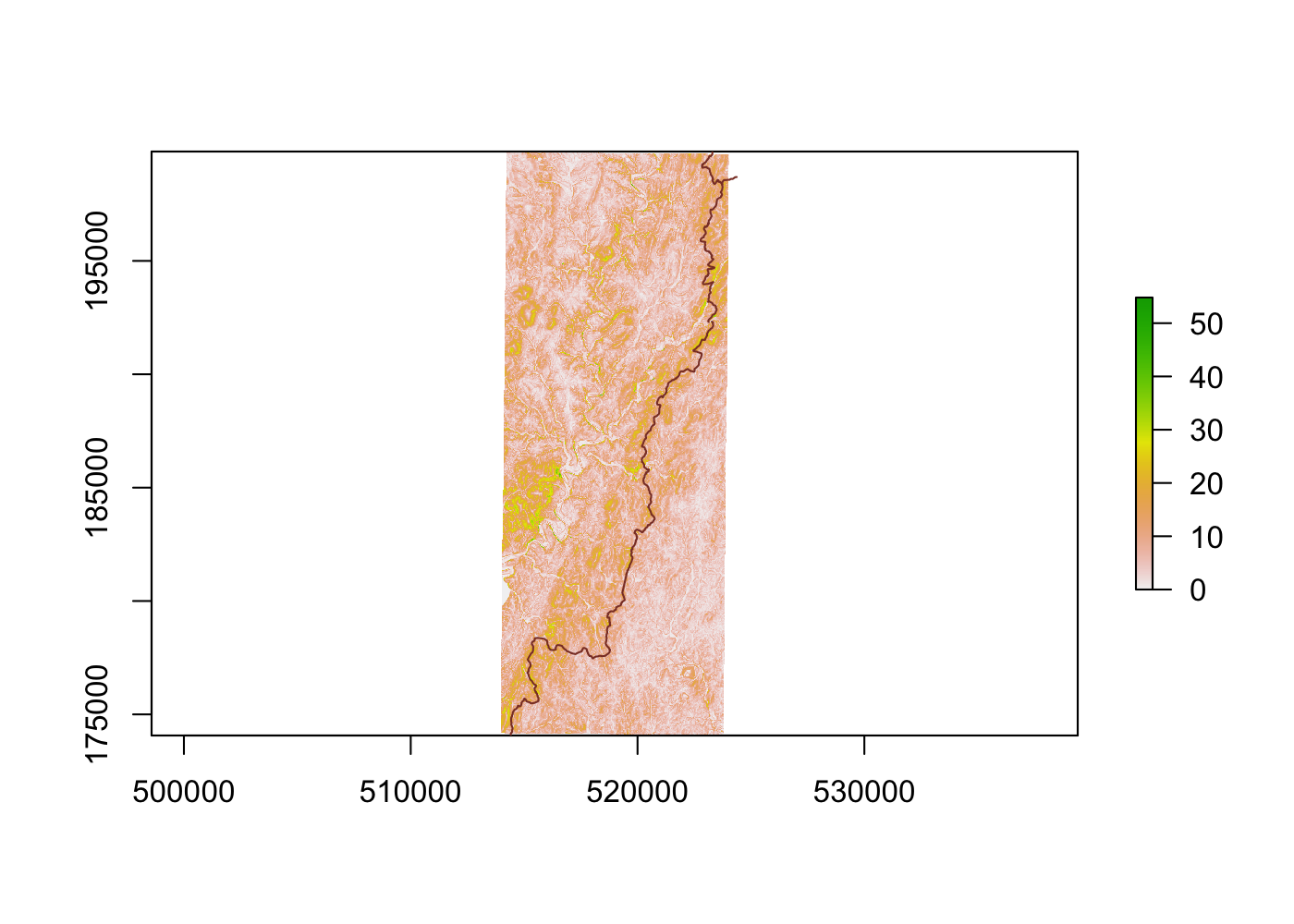
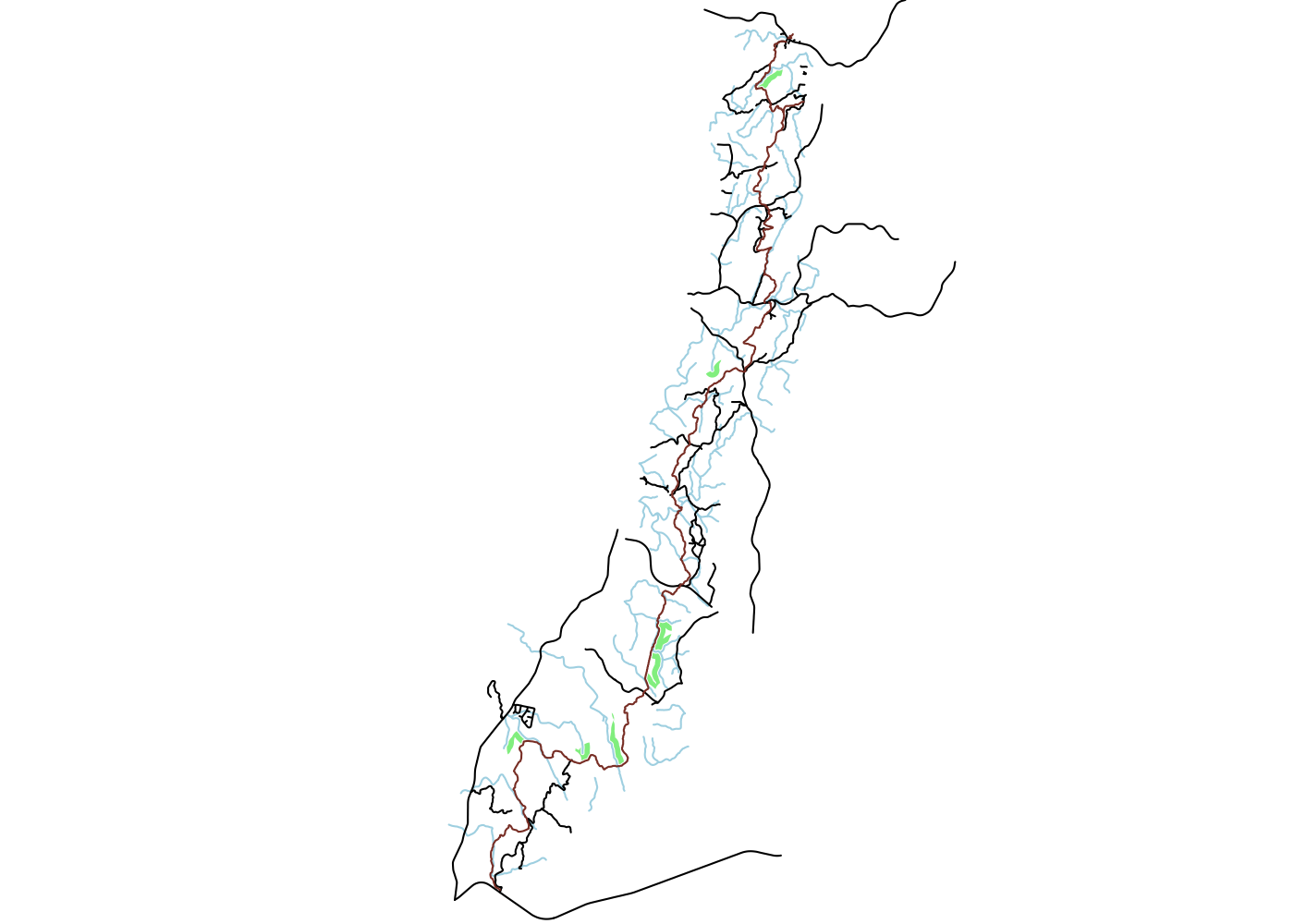

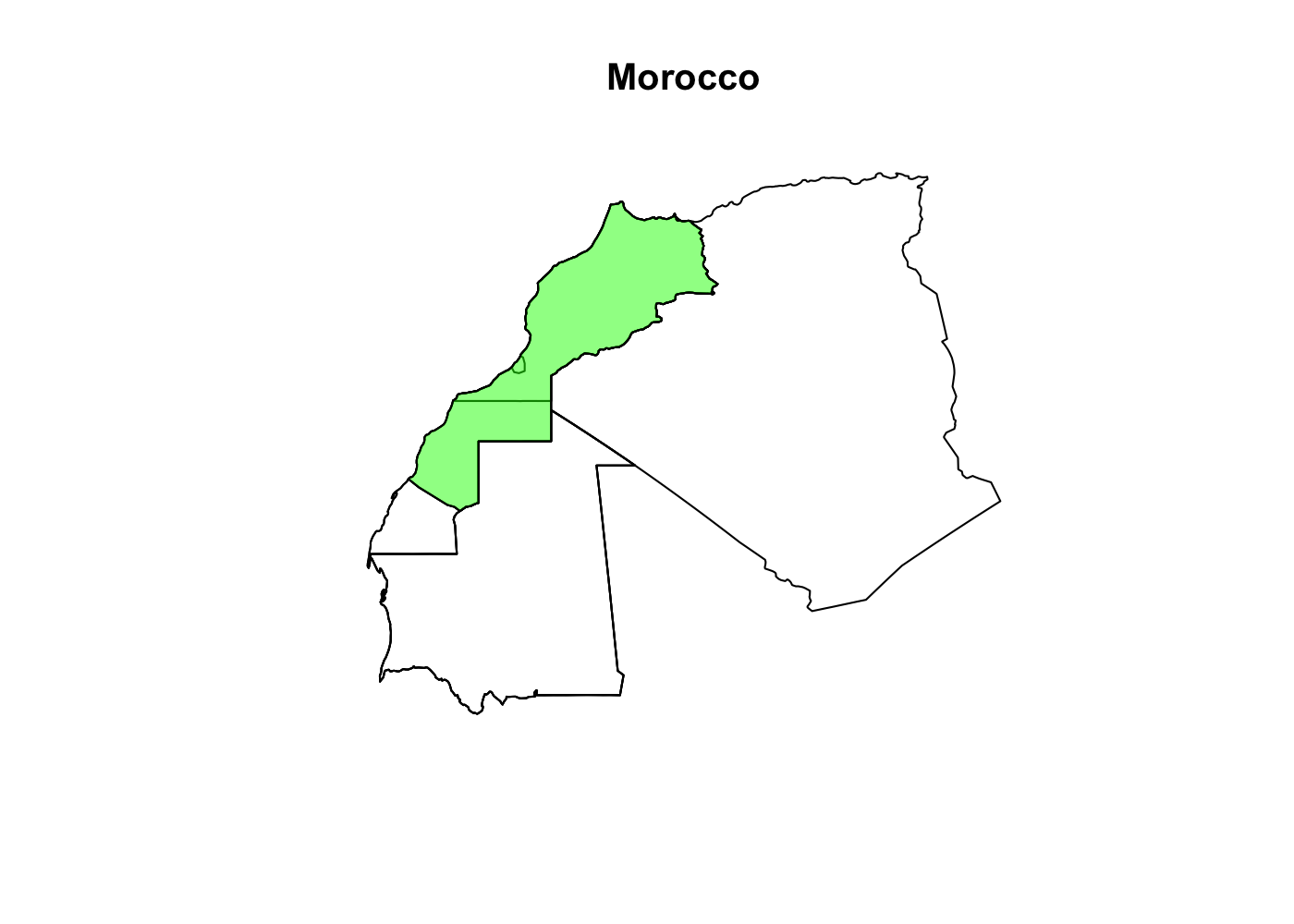
 Spatially filtering the GADM dataset is just as easy as with cshapes. To illustrate, I’m going to pull out a random polygon and use it to filter the data. However, these are third-order administrative divisions, and so it’s possible that even capturing all adjacent polygons won’t cover a very large area. To deal with this concern, we can buffer the polygon with the
Spatially filtering the GADM dataset is just as easy as with cshapes. To illustrate, I’m going to pull out a random polygon and use it to filter the data. However, these are third-order administrative divisions, and so it’s possible that even capturing all adjacent polygons won’t cover a very large area. To deal with this concern, we can buffer the polygon with the 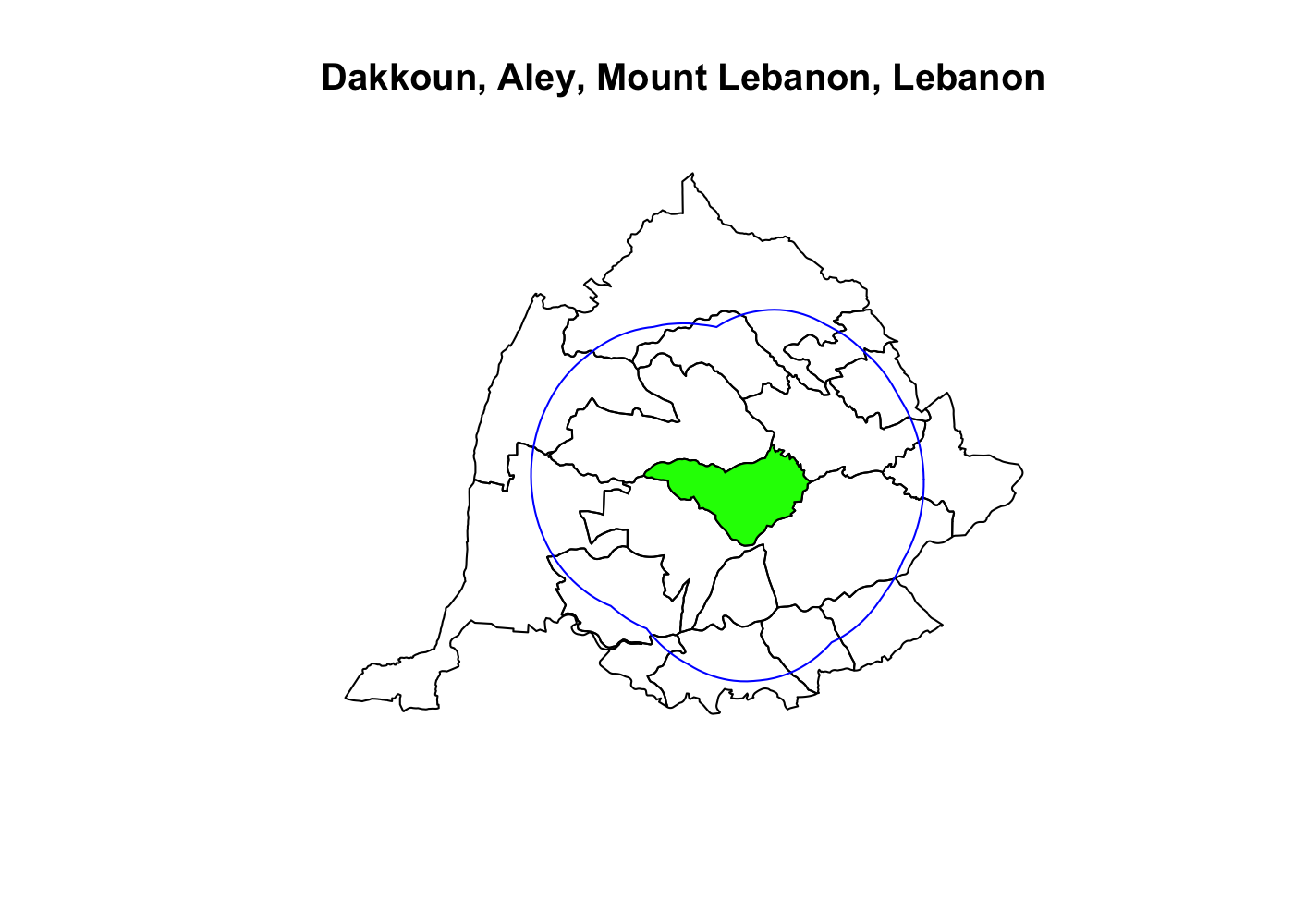
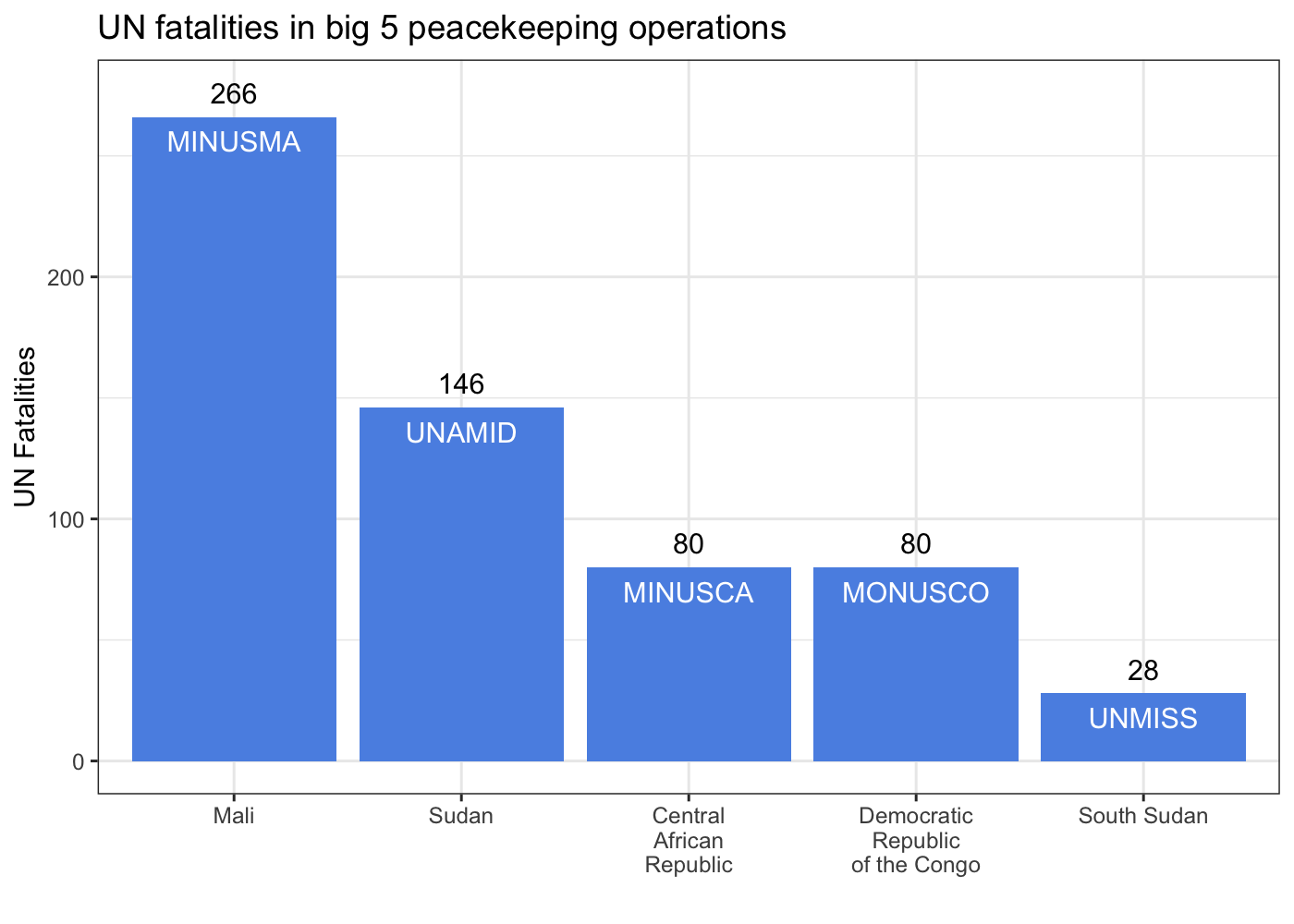
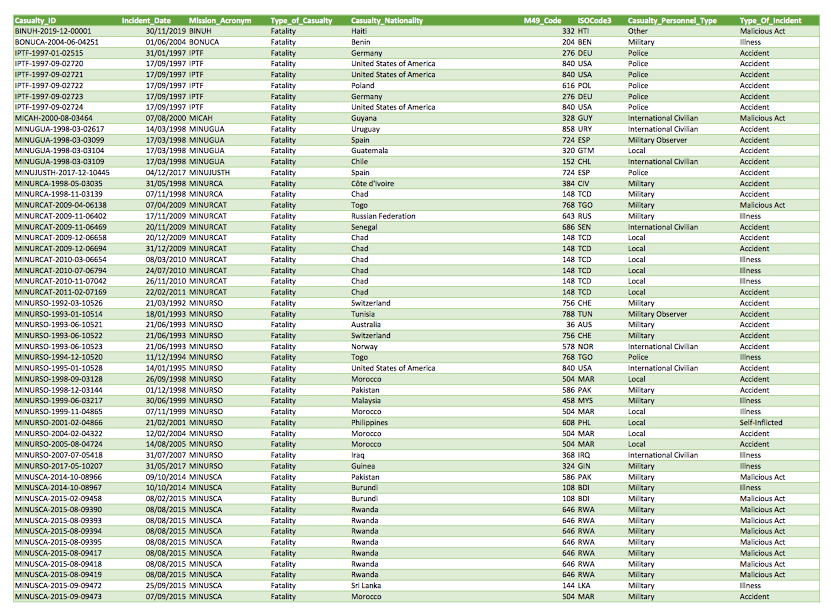 Since we were working on a short
deadline, I needed to get these data out of that PDF. The most direct
option is to just copy and paste the data into an Excel sheet. However,
these data run to 148 pages, so all that copying and pasting would be
tiring and risks introducing errors when your attention eventually slips
and you forget to include page 127.
Since we were working on a short
deadline, I needed to get these data out of that PDF. The most direct
option is to just copy and paste the data into an Excel sheet. However,
these data run to 148 pages, so all that copying and pasting would be
tiring and risks introducing errors when your attention eventually slips
and you forget to include page 127.